To me, saying that H3 is absent from the Near East, therefore it must be Paleolithic would be like saying that R1b-L21 is absent from the Near East, therefore it also must be Paleolithic. It doesn't really follow, does it? And mtDNA is very difficult to date accurately.
I admit that I didn't read the link. However the distribution speaks for itself. Both haplogroups are amazingly European, and extend as far as Russia and even Siberia, where R1a has settled, but where Middle Eastern farmers never went. Both haplogroups are too widespread geographically to be as recent as the Neolithic. I very seriously doubt that H* still existed in considerable numbers during the Neolithic. H first appeared circa 35,000 years, and started diversifying into today's top-level subclades from 22,000 to 12,000 years ago, i.e. before the Neolithic.
Ancient DNA tests showed that Haplogroup H17 might already have existed 25,000 years ago.
MtDNA is a very short sequence (compared to Y-DNA) and is particularly fickle when it comes to mutations. The same person can very well carry different mutations depending on what group of cells you test (in other words, take 3 times an MtDNA test and you
might get slightly different results each time, especially in the HVR1 and HVR2). New mtDNA mutations happen all the time inside the body. Some haplogroups seem to be more volatile than others in this regard (for example hg T and X). This is why I generally don't give much credit to studies of genetic diversity within mtDNA subclades.
Furthermore, the tree of haplogroup H is huge, new top-level subclades are still being found frequently (contrarily to more vertical haplogroups like the U's or K) and the structure of deep subclades has barely started to take form. At the last count on the
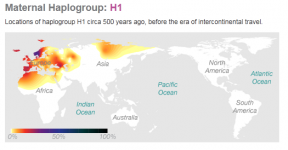
PhyloTree, haplogroup H1 had 31 first and second level subclades against only 9 for H3. 3 years ago there was almost nothing. In 3 years there might be hundreds for each (I wonder how they will proceed with the nomenclature for subclades after H1z). This is because it was very expensive until recently to test the full mtDNA sequence. The tree has really started to expand only 1 or 2 years ago.
Naturally there is always a bias at the beginning towards more heavily studied populations. If there was just a single detailed mtDNA study of the Maghreb and/or Iberia, this is enough to give the impression that this region has a greater genetic diversity than others. Jean Manco's post has four studies in reference: one of the Maghreb and three of Iberia. But nothing for other regions. How can they claim that H1 and H3 are more diverse in one region when they don't test other regions ? Besides, they only tested the HVRI and HVRII ! That's almost completely useless !
Yet, actually, it would make sense if Iberia (and the Caucasus) had a greater diversity of H1 and H3 than northern Europe for two reasons :
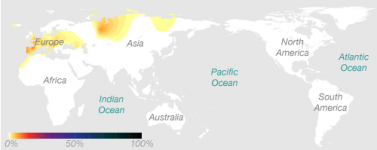
1) H1 and H3 both peak in Iberia in terms of percentages (more people means higher chances of mutations occurring)
2) Iberia served as one of the Ice Age refuges, from where European hunter-gatherers re-expanded. If H1 and H3 was confined to a few Ice Age refuges for a few thousand years, and southern European population were higher than northern European ones for many millennia after that, then it is to be expected that the genetic diversity be higher around southern European refuges. The North Caucasus is another peak for H1 and H3, and also served as an Ice Age refuge. The Balkans and Italy, which were the two other European refuges, don't have as much H1 and H3 simply because these regions were heavily settled by Neolithic farmers from the Near East.
Once again everything concurs in favour of a Paleolithic European origin of H1 and H3.