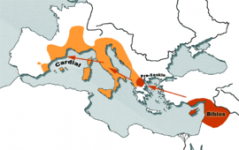
Finally a sizeable study of Neolithic Y-DNA in Western Europe !
R1b is indeed Indo-European, not linked to the diffusion of agriculture
We now have a definite confirmation that R1b was absent from Western Europe as far as the late Neolithic. The samples here only predate the supposed Indo-European invasion in the Bronze Age by circa 1000 years. The Neolithic site of Treilles is located between Narbonne and Perpignan, in the confine of the Languedoc and the Roussillon. Nowadays this region only has only about 5 or 6% of G2a and around 7% of I2a, against 60 to 70% of R1b1b2a1. This shows how quickly populations can change or haplogroups can be replaced.
European E1b1b and J could go back to the Greco-Roman Bronze and Iron Ages and later
The biggest surprise is the absence of haplogroups E1b1b and J. This was already the case in the German LBK site. This would imply that these two haplogroups expanded later. Dienekes suggested that E-V13 expanded from Greece during the Bronze Age. Although it might be true for the Balkans region and Greek colonies, I cannot imagine how the Greeks would be responsible for the presence of haplogroup E in as far as Portugal, Britain, the Benelux, Germany, Poland or Belarus.
I would imagine that both E1b1b and J came from the Near East to Greece and Italy during the Bronze Age, and founded the Minoan and Etruscan civilizations there. Later, the Roman Empire would have allowed people from Greece, Italy and the whole Near and Middle East to travel to Western and Central Europe and spread those lineages. Haplogroups would have mixed progressively across Europe during the Roman period, and continued during the Middle Ages and in modern times. With each century that passes the population of European cities is getting increasingly homogeneous in terms of haplogroup percentages, due to the continuous movement of people.
I have long wondered how it was possible that Iceland had a near complete absence of Near Eastern haplogroups (G2a, E1b1b, J and T) whereas the rest of Scandinavia has a small but significant percentage (ranging from 6.5% in Denmark to 2.5% Sweden). Since Icelandic people hailed from Norway and Denmark, the absence of E, G, J and T in modern Icelandic means that these haplogroups has not yet reached Scandinavia when Iceland was settled 1000 years ago.
Neolithic MtDNA
In the
supplemental data, I counted the following mitochondrial haplogroups among the 29 samples :
- U => 1 individual
- U5 => 4 individuals
- U5b1c => 1 individual
- K1a => 2 individuals
- HV0 => 2 individuals
- H1 => 3 individuals
- H3 => 3 individuals
- V => 1 individual
- J1 => 6 individuals
- T2b => 2 individuals
- X2 => 4 individuals
Among these, I would place the U5, H1, H3 and V as being of Paleolithic European origin (because they are rare or absent from the Middle East), and the rest as Neolithic migrants from the Near/Middle East. This would give us 41% of Paleolithic maternal lineages. It is to be expected that early farmers married girls from the local hunter-gatherer community, rather than the other way round.
Note the high percentage of X2 (13.8%) which is fairly rare nowadays and is strongly associated with the Caucasus. I have always thought of X2 as the maternal equivalent of G2a, which is why I used the same grey colour for both haplogroups on this website.
K1a, J1 and T2b are also somewhat typical of the greater Caucasus region, including Anatolia the Pontic steppes. These haplogroups are found as well among Middle Easterners as in Eurasian steppe populations, most certainly due to the exchange of wives across the Caucasus region.