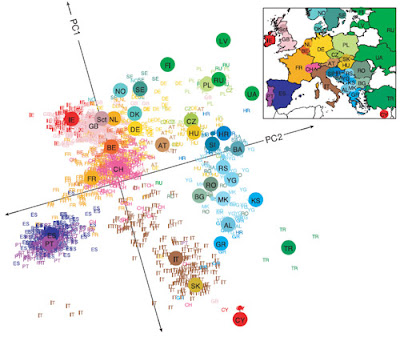
I have made distribution maps for most of the major haplogroups. I thought it would be interesting to compare Y-DNA distributions with autosomal admixtures. I used the data for the Mediterranean admixture from the Dodecad Project to make this map. It's far from perfect because many key populations were missing (Croatians, Bosnians, Serbs, Czechoslovaks, Belarusians, Ukrainians, most of the Caucasus beside Georgia and Armenia). I used the data from individual members to fill the gaps.
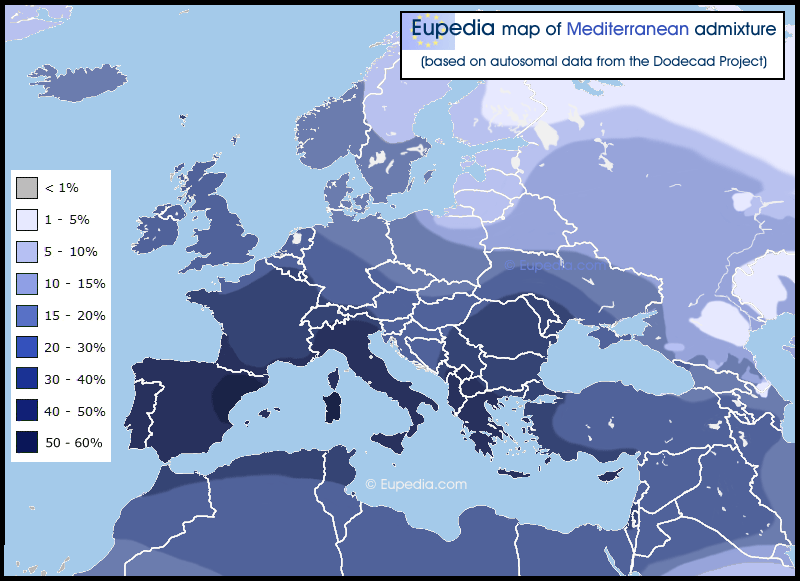
In my opinion, this Mediterranean admixture has a strong E1b1b (E-M78 and subclades, though not E-M81) component to it, as well as a likely I2a1 influence in places like Northeast Spain and Sardinia. I believe that R1b in Western Europe is hiding a lot of anterior lineages, which could have been principally I2a and E-M78.
Autosomal genes are naturally more evenly spread out inside a population sharing the same borders or language. That is because women are more likely to be "imported" as wives from other regions, and also because autosomal genes mix very quickly in the gene pool. A third reason the autosomal map looks more evenly distributed is that I didn't have as much data for regional populations (mostly country averages).
There is no correspondence with G2a, J2 or J1 because this Mediterranean admixture is lowest in the Mediterranean region where G2a, J1 and J2 are the highest.
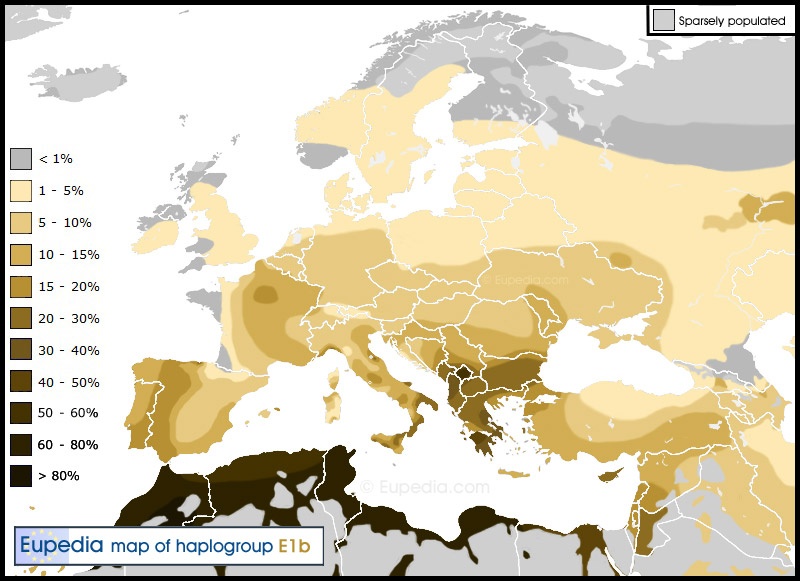
I doubt that it is a coincidence that this Mediterranean admixture matches quite well the so-called Mediterranean Race described by anthropologist Madison Grant. One of the main characteristics of Mediterranean people is the dolichocephalic skull shape (long head), which they have in common with the Nordics.


In my opinion, this Mediterranean admixture has a strong E1b1b (E-M78 and subclades, though not E-M81) component to it, as well as a likely I2a1 influence in places like Northeast Spain and Sardinia. I believe that R1b in Western Europe is hiding a lot of anterior lineages, which could have been principally I2a and E-M78.
Autosomal genes are naturally more evenly spread out inside a population sharing the same borders or language. That is because women are more likely to be "imported" as wives from other regions, and also because autosomal genes mix very quickly in the gene pool. A third reason the autosomal map looks more evenly distributed is that I didn't have as much data for regional populations (mostly country averages).
There is no correspondence with G2a, J2 or J1 because this Mediterranean admixture is lowest in the Mediterranean region where G2a, J1 and J2 are the highest.

I doubt that it is a coincidence that this Mediterranean admixture matches quite well the so-called Mediterranean Race described by anthropologist Madison Grant. One of the main characteristics of Mediterranean people is the dolichocephalic skull shape (long head), which they have in common with the Nordics.