Angela
Elite member
- Messages
- 21,823
- Reaction score
- 12,329
- Points
- 113
- Ethnic group
- Italian
This promises to be very interesting. It was just presented at a conference, so all we have is the abstract (thanks to anthrogenica), but hopefully the paper will be out soon. It seems it's just about as predicted.
"Fine-Scale Human Genetic Structure in France
Aude Saint-Pierre, Céline Bellenguez, Luc Letenneur, Claudine Berr, Carole Dufouil, Philippe Amouyel,Emmanuelle Génin
Characterizing geographical population structure is critical to genetic studies of disease as it is an important cause of false positive results in genome wide association studies (GWASs). The genetic structure of several countries in Europe has been carefully studied but there is a lack of descriptive study of the French population. Indeed, apart from the work of Karakachoff et al (2015) that focused on the western part of France and detected interesting stratification, no study so far provided a comprehensive look at the French genetic landscape. Here we describe the genetic structure of the French population at a fine-scale using genetic data from the 3 Cities (3C) Study, a population cohort of French elderly individuals that served as controls in several GWAS conducted with French patients. From this cohort, we had access to 4,433 genotyped individuals sampled in three regions of France but born all over France.
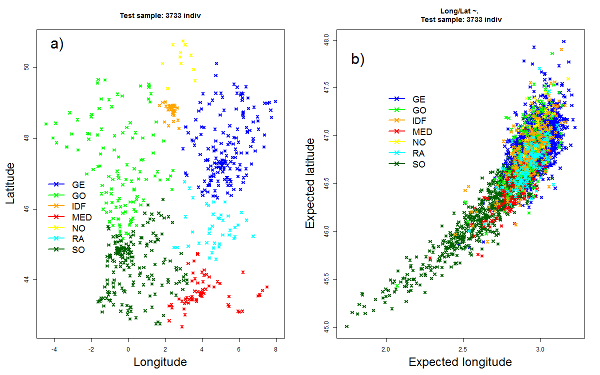
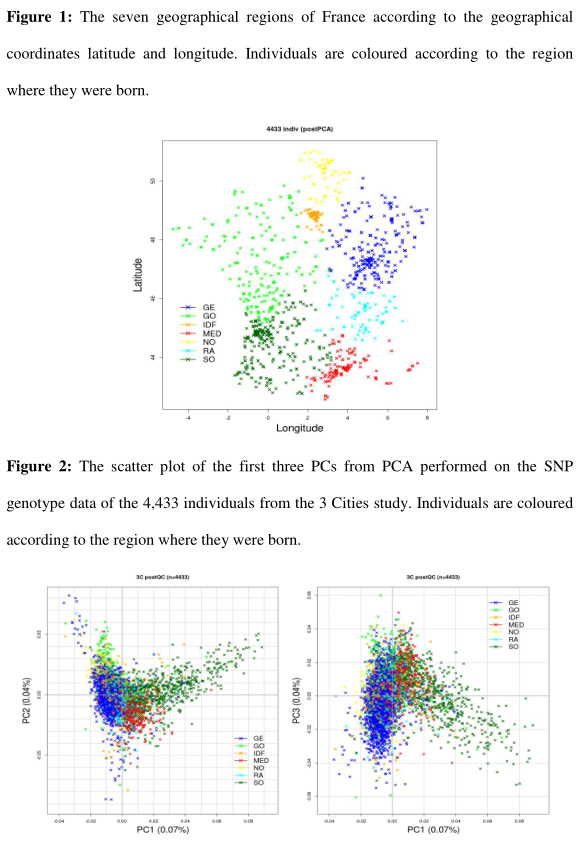
We selected a subset of 770 individuals to cover evenly the different regions of France and applied methods that utilize haplotype information for detecting fine-scale population structure. The 770 individuals were partitioned into homogeneous clusters using CHROMOPAINTER and fineSTRUCTURE analysis (Lawson et al. 2012). Six clusters were identified that correlate well with the geographic origin of individuals. The coarsest level of genetic differentiation separates the samples from southwestern French from all the others. Subsequent splits reveal more subtle differentiation except for samples from western France which showed a relatively high degree of homogeneity.
For each cluster we used CHROMOPAINTER to estimate an “ancestry profile” which characterises the ancestry of the cluster as a mixture of the reference sample. Using the subsample of 770 individuals as a reference sample to assign the remaining 3C individuals, we found that the cluster assignment was coherent with the places of birth of individuals. The same procedure was applied using the five European samples from the 1000 Genomes Project as a reference sample. Contribution from European populations shows a cline roughly north-south, in ancestry profiles. Spain (IBS) is the largest contributor of the southwest and south clusters while the highest contribution of Great-Britain (GBR) population is observed in Brittany.
In conclusion, we provide evidence that there exist some levels of genetic stratification in France. The French population could roughly be divided into 6 genetic clusters that correlate well with geography. The knowledge of this stratification pattern will be useful to design robust and powerful association studies."
"Fine-Scale Human Genetic Structure in France
Aude Saint-Pierre, Céline Bellenguez, Luc Letenneur, Claudine Berr, Carole Dufouil, Philippe Amouyel,Emmanuelle Génin
Characterizing geographical population structure is critical to genetic studies of disease as it is an important cause of false positive results in genome wide association studies (GWASs). The genetic structure of several countries in Europe has been carefully studied but there is a lack of descriptive study of the French population. Indeed, apart from the work of Karakachoff et al (2015) that focused on the western part of France and detected interesting stratification, no study so far provided a comprehensive look at the French genetic landscape. Here we describe the genetic structure of the French population at a fine-scale using genetic data from the 3 Cities (3C) Study, a population cohort of French elderly individuals that served as controls in several GWAS conducted with French patients. From this cohort, we had access to 4,433 genotyped individuals sampled in three regions of France but born all over France.
We selected a subset of 770 individuals to cover evenly the different regions of France and applied methods that utilize haplotype information for detecting fine-scale population structure. The 770 individuals were partitioned into homogeneous clusters using CHROMOPAINTER and fineSTRUCTURE analysis (Lawson et al. 2012). Six clusters were identified that correlate well with the geographic origin of individuals. The coarsest level of genetic differentiation separates the samples from southwestern French from all the others. Subsequent splits reveal more subtle differentiation except for samples from western France which showed a relatively high degree of homogeneity.
For each cluster we used CHROMOPAINTER to estimate an “ancestry profile” which characterises the ancestry of the cluster as a mixture of the reference sample. Using the subsample of 770 individuals as a reference sample to assign the remaining 3C individuals, we found that the cluster assignment was coherent with the places of birth of individuals. The same procedure was applied using the five European samples from the 1000 Genomes Project as a reference sample. Contribution from European populations shows a cline roughly north-south, in ancestry profiles. Spain (IBS) is the largest contributor of the southwest and south clusters while the highest contribution of Great-Britain (GBR) population is observed in Brittany.
In conclusion, we provide evidence that there exist some levels of genetic stratification in France. The French population could roughly be divided into 6 genetic clusters that correlate well with geography. The knowledge of this stratification pattern will be useful to design robust and powerful association studies."