Angela
Elite member
- Messages
- 21,823
- Reaction score
- 12,329
- Points
- 113
- Ethnic group
- Italian
[h=1]Genetic Heterogeneity in Algerian Human Populations[/h]This is the link to the paper:
http://journals.plos.org/plosone/article?id=10.1371/journal.pone.0138453#pone.0138453.s010
"Our results of uniparental and autosomal markers in Algeria agree with the presence of ancestral components previously described in North Africa, attesting the genome complexity of Algerians (S8 Table). Concerning the Y-chromosome data, the highest frequencies are seen for the autochthonous North African lineages E-M81 and E-M78, this last one more frequent in Northeastern Africa where it has probably emerged [39]; whereas the presence of the Middle Eastern Y-chromosome J1-M267 has been attributed to the Islamic expansion [40]. In a similar way, the mitochondrial DNA analysis shows also different lineages in Algeria, already observed in North Africa: the North African lineages U6 and M1 that have been dated to Paleolithic times [41–43], the Eurasian H (related to the Neolithic expansion) and HV [29,44,45]; and the sub-Saharan lineages (L). It has been suggested that the sub-Saharan lineages for both mtDNA and Y-chromosome reached very recently North Africa through the slave trade routes across the Sahara [16,46,47]. Moreover, the autosomal genome-wide SNPs analysis also demonstrates the admixture of the Eurasian and African components in both Berber (Mozabite and Zenata) and non-Berber populations from Algeria in agreement with the general genetic North African landscape [3].
The genetic structure observed in the Algerian analyzed groups is neither fully correlated with ethnic affiliation (Berber-Arab) nor with geography (coast vs. inland). It is noteworthy, however, that when we removed the Reguibate population from the Arab group, a significant differentiation was observed between the paternal gene pool profiles of Arabs and Berbers. This absence of differentiation between Arabs and Berbers is in agreement with what has been already observed in several North African populations by the analysis of different genetic markers, such as autosomal classical markers (such as HLA markers and GM allotypes) [6,12,14]; autosomal STRs [7]; Alu polymorphisms [8]; Y-chromosome [13,48]; and mtDNA analyses [11,12]. As a result, it has been suggested that the Arabization of North African populations was mainly a cultural process rather than a demographic replacement of autochthonous groups [11]."
This is also interesting:
"The distribution and frequencies of the North African, Eurasian and sub-Saharan components both in uniparental and autosomal markers is variable in each group, not only when comparing Berber and non-Berber, but also within linguistic groups. For example, some autochthonous North African haplogroups were not present in certain samples, such as the mtDNA U6 haplogroup that was absent in Algiers and present in only one individual in the analyzed Zenata group. In the same way, mtDNA haplogroup M1 is absent in the Zenata population. The absence of the maternal North African component in these groups, especially the Zenata Berbers, might be explained by extensive genetic drift and the remarkable high frequency of sub-Saharan lineages (~23% for the Y-chromosome E-M2 haplogroup and ~ 65% of mtDNA L lineages) in the Zenata sample. Our autosomal analysis also shows the close position of the Zenata group to the sub-Saharan populations, and the high variance in this sub-Saharan ancestry suggest that this group has experienced recent gene flow."
This is in accordance with comments that have been made on this site:
" The Berbers of North Africa are those North Africans who have retained at least a partial use of the Berber language and some degree of their tribal structure and their ancient culture. They are usually pretty endogamous. The level of SSA among them varies depending on the tribe and the country. Some, like the Mozabites, have quite a bit. Other groups look like they have a lot less, and some among them just look southern European.Likewise there are a lot of North Africans who don't primarily identify as Berbers. They also vary in terms of SSA admixture, with some urban, upper class people not showing a lot of SSA but most of them showing what is close to what the studies say, which is about 20-25% SSA ancestry."
The "SSA" like ancestry that this paper reports for some Berber groups is very high, whereas for other Berber groups like the Mozabites it is much lower, to the best of my recollection, than was found in prior analyses, where it is reported as about 20% in some studies. It may be because they use only the Yoruba as a reference population, and do not include some East African groups. The same effect is in operation in terms of the Palestinians, who get 10-12% SSA in some studies, but much lower here.
This is the table:
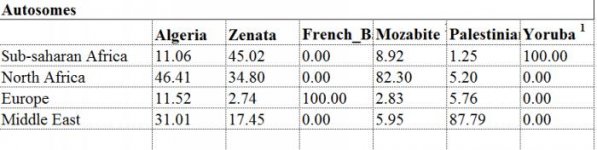
http://journals.plos.org/plosone/article?id=10.1371/journal.pone.0138453#pone.0138453.s010
"Our results of uniparental and autosomal markers in Algeria agree with the presence of ancestral components previously described in North Africa, attesting the genome complexity of Algerians (S8 Table). Concerning the Y-chromosome data, the highest frequencies are seen for the autochthonous North African lineages E-M81 and E-M78, this last one more frequent in Northeastern Africa where it has probably emerged [39]; whereas the presence of the Middle Eastern Y-chromosome J1-M267 has been attributed to the Islamic expansion [40]. In a similar way, the mitochondrial DNA analysis shows also different lineages in Algeria, already observed in North Africa: the North African lineages U6 and M1 that have been dated to Paleolithic times [41–43], the Eurasian H (related to the Neolithic expansion) and HV [29,44,45]; and the sub-Saharan lineages (L). It has been suggested that the sub-Saharan lineages for both mtDNA and Y-chromosome reached very recently North Africa through the slave trade routes across the Sahara [16,46,47]. Moreover, the autosomal genome-wide SNPs analysis also demonstrates the admixture of the Eurasian and African components in both Berber (Mozabite and Zenata) and non-Berber populations from Algeria in agreement with the general genetic North African landscape [3].
The genetic structure observed in the Algerian analyzed groups is neither fully correlated with ethnic affiliation (Berber-Arab) nor with geography (coast vs. inland). It is noteworthy, however, that when we removed the Reguibate population from the Arab group, a significant differentiation was observed between the paternal gene pool profiles of Arabs and Berbers. This absence of differentiation between Arabs and Berbers is in agreement with what has been already observed in several North African populations by the analysis of different genetic markers, such as autosomal classical markers (such as HLA markers and GM allotypes) [6,12,14]; autosomal STRs [7]; Alu polymorphisms [8]; Y-chromosome [13,48]; and mtDNA analyses [11,12]. As a result, it has been suggested that the Arabization of North African populations was mainly a cultural process rather than a demographic replacement of autochthonous groups [11]."
This is also interesting:
"The distribution and frequencies of the North African, Eurasian and sub-Saharan components both in uniparental and autosomal markers is variable in each group, not only when comparing Berber and non-Berber, but also within linguistic groups. For example, some autochthonous North African haplogroups were not present in certain samples, such as the mtDNA U6 haplogroup that was absent in Algiers and present in only one individual in the analyzed Zenata group. In the same way, mtDNA haplogroup M1 is absent in the Zenata population. The absence of the maternal North African component in these groups, especially the Zenata Berbers, might be explained by extensive genetic drift and the remarkable high frequency of sub-Saharan lineages (~23% for the Y-chromosome E-M2 haplogroup and ~ 65% of mtDNA L lineages) in the Zenata sample. Our autosomal analysis also shows the close position of the Zenata group to the sub-Saharan populations, and the high variance in this sub-Saharan ancestry suggest that this group has experienced recent gene flow."
This is in accordance with comments that have been made on this site:
" The Berbers of North Africa are those North Africans who have retained at least a partial use of the Berber language and some degree of their tribal structure and their ancient culture. They are usually pretty endogamous. The level of SSA among them varies depending on the tribe and the country. Some, like the Mozabites, have quite a bit. Other groups look like they have a lot less, and some among them just look southern European.Likewise there are a lot of North Africans who don't primarily identify as Berbers. They also vary in terms of SSA admixture, with some urban, upper class people not showing a lot of SSA but most of them showing what is close to what the studies say, which is about 20-25% SSA ancestry."
The "SSA" like ancestry that this paper reports for some Berber groups is very high, whereas for other Berber groups like the Mozabites it is much lower, to the best of my recollection, than was found in prior analyses, where it is reported as about 20% in some studies. It may be because they use only the Yoruba as a reference population, and do not include some East African groups. The same effect is in operation in terms of the Palestinians, who get 10-12% SSA in some studies, but much lower here.
This is the table: