Angela
Elite member
- Messages
- 21,823
- Reaction score
- 12,329
- Points
- 113
- Ethnic group
- Italian
Human adaptation and population differentiation in the light of ancient genomes
See:
http://www.nature.com/ncomms/2016/160318/ncomms10775/full/ncomms10775.html
The supplement:
http://www.nature.com/ncomms/2016/160318/ncomms10775/full/ncomms10775.html#supplementary-information
"The influence of positive selection sweeps in human evolution is increasingly debated, although our ability to detect them is hampered by inherent uncertainties in the timing of past events. Ancient genomes provide snapshots of allele frequencies in the past and can help address this question. We combine modern and ancient genomic data in a simple statistic (DAnc) to time allele frequency changes, and investigate the role of drift and adaptation in population differentiation. Only 30% of the most strongly differentiated alleles between Africans and Eurasians changed in frequency during the colonization of Eurasia, but in Europe these alleles are enriched in genic and putatively functional alleles to an extent only compatible with local adaptation. Adaptive alleles—especially those associated with pigmentation—are mostly of hunter-gatherer origin, although lactose persistence arose in a haplotype present in farmers. These results provide evidence for a role of local adaptation in human population differentiation."
I've only skimmed the paper and the supplement, but from what I can see their argument is not based on finding the linked snps to these adaptations in any of the ancient samples, but to finding background haplotypes that might have given rise to these snps.
The ancient genomes they examined were Loschbour, Stuttgart, and Ust-Ishim.
It certainly makes sense that a background haplotype for digestion of animal milk might be present in farmers, and that one for lighter skin might be present in people from more northern latitudes (SLC45A2). That doesn't explain why the actual derived version is present in Anatolian farmers, if not specifically in Stuttgart.
There's also the East Asian light skin alleles to explain. The authors maintain that they can't pick up a signal of selection for them because there has been more drift among East Asians. I have to reread that section.
See:
http://www.nature.com/ncomms/2016/160318/ncomms10775/full/ncomms10775.html
The supplement:
http://www.nature.com/ncomms/2016/160318/ncomms10775/full/ncomms10775.html#supplementary-information
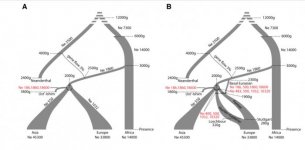
"The influence of positive selection sweeps in human evolution is increasingly debated, although our ability to detect them is hampered by inherent uncertainties in the timing of past events. Ancient genomes provide snapshots of allele frequencies in the past and can help address this question. We combine modern and ancient genomic data in a simple statistic (DAnc) to time allele frequency changes, and investigate the role of drift and adaptation in population differentiation. Only 30% of the most strongly differentiated alleles between Africans and Eurasians changed in frequency during the colonization of Eurasia, but in Europe these alleles are enriched in genic and putatively functional alleles to an extent only compatible with local adaptation. Adaptive alleles—especially those associated with pigmentation—are mostly of hunter-gatherer origin, although lactose persistence arose in a haplotype present in farmers. These results provide evidence for a role of local adaptation in human population differentiation."
I've only skimmed the paper and the supplement, but from what I can see their argument is not based on finding the linked snps to these adaptations in any of the ancient samples, but to finding background haplotypes that might have given rise to these snps.
The ancient genomes they examined were Loschbour, Stuttgart, and Ust-Ishim.
It certainly makes sense that a background haplotype for digestion of animal milk might be present in farmers, and that one for lighter skin might be present in people from more northern latitudes (SLC45A2). That doesn't explain why the actual derived version is present in Anatolian farmers, if not specifically in Stuttgart.
There's also the East Asian light skin alleles to explain. The authors maintain that they can't pick up a signal of selection for them because there has been more drift among East Asians. I have to reread that section.