Tomenable
Regular Member
- Messages
- 5,419
- Reaction score
- 1,337
- Points
- 113
- Location
- Poland
- Ethnic group
- Polish
- Y-DNA haplogroup
- R1b-L617
- mtDNA haplogroup
- W6a
Post your Eurogenes Steppe K10 results (if you tried this calculator).
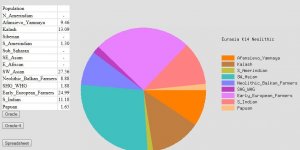
I have a very high % of Hindu Kush (6-7 times higher than Polish average from spreadsheet):
Polish average includes 11 samples: 10H, 11H, 12H, 13H, 14H, 15H, 16H, 6H, 7H, 8H, 9H.
Here is the spreadsheet: https://docs.google.com/spreadsheet...cmhWtsYv0p39avjqM-G3-6Xew/edit#gid=1809893991
How To:
It’s not complicated. You guys can do it very quickly. Let’s try a last time.
So forget the “readme”. Just follow the steps below.
Let’s presume that LeBrok tested in 23andMe, and that Fluffy tested in FTDNA. LeBrok will download his 23andMe raw data and name the file as LeBrok.txt , while Fluffy will download his FTDNA raw data (Build 37 Raw Data Concatenated) and name the file as Fluffy.csv . Both using Windows.
Download SteppeK10.zip (save icon - an arrow in top right): https://drive.google.com/file/d/0B-X...lRSkcwV0k/view
Extract the files into a folder called SteppeK10, in C:/ . Now you have the folder C:/SteppeK10 , right? Inside it, the files DIYDodecadWin.exe , standardize.r , Steppe.10.F , Steppe.alleles , steppe.par , Steppe.txt .
LeBrok and Fluffy will put the files LeBrok.txt and Fluffy.csv into the folder C:/SteppeK10, respectively.
Download R software: http://cran.utstat.utoronto.ca/bin/w...-3.3.1-win.exe
Install* this program in English and using default options (if you don’t know how to do it, just ask). After that, run it (click in the icon “R x.. 3.3.1” in Desktop).
*I installed just the 64 bits version.
Once R is running, click in “File” (top left), then click in “Change dir”. Select the folder C:/SteppeK10 and click in “ok”.
Do you see the R console? After the “>” character, type the command below and press Enter:
source('standardize.r')
Now:
a. LeBrok, who tested hypothetically with 23andMe, will type the command below, press Enter and wait til he can type anything again:
standardize('LeBrok.txt', company='23andMe')
b. Fluffy will do that with this command:
standardize('Fluffy.csv', company='ftdna')
Finally, you both will type the command below and wait results show up:
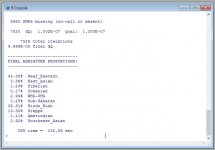
system('DIYDodecadWin steppe.par')
I have a very high % of Hindu Kush (6-7 times higher than Polish average from spreadsheet):
| Admixture: | Tomenable: | Average of 11 Poles: |
| WHG-UHG | 37.14 | 39.59 |
| Steppe | 29.54 | 33.87 |
| Near Eastern | 29.36 | 24.41 |
| Hindu Kush | 3.32 | 0.50 |
| Oceanian | 0.09 | 0.18 |
| Siberian | 0.05 | 0.62 |
| East Asian | - | 0.16 |
| Amerindian | 0.50 | 0.34 |
| Southeast Asian | - | 0.33 |
| Sub-Saharan | - | - |
Polish average includes 11 samples: 10H, 11H, 12H, 13H, 14H, 15H, 16H, 6H, 7H, 8H, 9H.
Here is the spreadsheet: https://docs.google.com/spreadsheet...cmhWtsYv0p39avjqM-G3-6Xew/edit#gid=1809893991
How To:
It’s not complicated. You guys can do it very quickly. Let’s try a last time.
So forget the “readme”. Just follow the steps below.
Let’s presume that LeBrok tested in 23andMe, and that Fluffy tested in FTDNA. LeBrok will download his 23andMe raw data and name the file as LeBrok.txt , while Fluffy will download his FTDNA raw data (Build 37 Raw Data Concatenated) and name the file as Fluffy.csv . Both using Windows.
Download SteppeK10.zip (save icon - an arrow in top right): https://drive.google.com/file/d/0B-X...lRSkcwV0k/view
Extract the files into a folder called SteppeK10, in C:/ . Now you have the folder C:/SteppeK10 , right? Inside it, the files DIYDodecadWin.exe , standardize.r , Steppe.10.F , Steppe.alleles , steppe.par , Steppe.txt .
LeBrok and Fluffy will put the files LeBrok.txt and Fluffy.csv into the folder C:/SteppeK10, respectively.
Download R software: http://cran.utstat.utoronto.ca/bin/w...-3.3.1-win.exe
Install* this program in English and using default options (if you don’t know how to do it, just ask). After that, run it (click in the icon “R x.. 3.3.1” in Desktop).
*I installed just the 64 bits version.
Once R is running, click in “File” (top left), then click in “Change dir”. Select the folder C:/SteppeK10 and click in “ok”.
Do you see the R console? After the “>” character, type the command below and press Enter:
source('standardize.r')
Now:
a. LeBrok, who tested hypothetically with 23andMe, will type the command below, press Enter and wait til he can type anything again:
standardize('LeBrok.txt', company='23andMe')
b. Fluffy will do that with this command:
standardize('Fluffy.csv', company='ftdna')
Finally, you both will type the command below and wait results show up:
system('DIYDodecadWin steppe.par')
Last edited by a moderator: