Install the app
How to install the app on iOS
Follow along with the video below to see how to install our site as a web app on your home screen.

Note: This feature currently requires accessing the site using the built-in Safari browser.
You are using an out of date browser. It may not display this or other websites correctly.
You should upgrade or use an alternative browser.
You should upgrade or use an alternative browser.
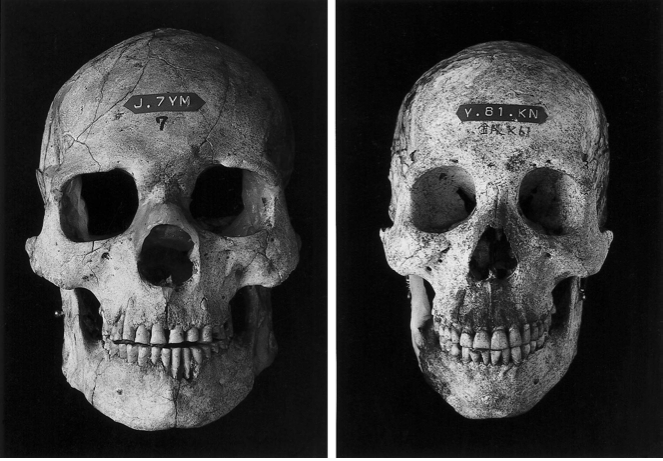
classify Amazon and Jomon skull
- Thread starter johen
- Start date
ThirdTerm
Regular Member
- Messages
- 187
- Reaction score
- 97
- Points
- 28
Two Cro-Magnons' mtDNA haplogroup (23,000-year-old Paglicci 52 and 24,720-year-old Paglicci 12) was identified as haplogroup N and mtDNA haplogroups N9b and M7a are called the Jomon haplotype. More specifically, the two Cro-Magnons carried macrohaplogroup N, containing haplogroups W, X, I, N1a, N1b, N1c, and N* (Caramelli et al. 2003). The Jomon and Cro-Magnons share common maternal ancestry, which may have contributed to the morphological similarity between them. Moreover, Kennewick Man's mtDNA haplogroup was recently identified as haplogroup X2a by Rasmussen et al. (2015) and some Native Americans with this haplotype are distantly related to Cro-Magnons.
Specific mtDNA sites outside HVRI were also analyzed (by amplification, cloning, and sequencing of the surrounding region) to classify more precisely the ancient sequences within the phylogenetic network of present-time mtDNAs (35, 36). Paglicci-25 has the following motifs: +7,025 AluI, 00073A, 11719G, and 12308A. Therefore, this sequence belongs to either haplogroups HV or pre-HV, two haplogroups rare in general but with a comparatively high frequencies among today's Near-Easterners (35). Paglicci-12 shows the motifs 00073G, 10873C, 10238T, and AACC between nucleotide positions 10397 and 10400, which allows the classification of this sequence into the macrohaplogroupN,containing haplogroups W, X, I, N1a, N1b, N1c, and N*. Following the definition given in ref. 36, the presence of a single mutation in 16,223 within HRVI suggests a classification of Paglicci-12 into the haplogroup N*, which is observed today in several samples from the Near East and, at lower frequencies, in the Caucasus (35). It is difficult to say whether the apparent evolutionary relationship between Paglicci-25 and Paglicci-12 and those populations is more than a coincidence. Indeed, the haplogroups to which the Cro-Magnon type sequences appear to belong are rare among modern samples, and therefore their frequencies are poorly estimated. However, genetic affinities between the first anatomically modern Europeans and current populations of the Near East make sense in the light of the likely routes of Upper Paleolithic human expansions in Europe, as documented in the archaeological record (37).
Last edited:
This thread has been viewed 3874 times.