@FrankN,
I don't think this is quite the thread for a detailed discussion of Lazaridis et al, and I'm very pressed for time as I'm about to go on “sabbatical” for about a six to eight week period, but I don’t want it to appear as if I am ignoring your post, and, frankly, I am loathe to let what I consider serious misunderstandings of the paper stand for those who may not be very familiar with autosomal analysis.
(I want to extend my apologies to Aberdeen and the moderators for continuing this hijacking of this thread. Please feel free to remove both of these extended posts about Lazaridis et al to a more appropriate venue.)
So, to begin, you stated that I misrepresented the findings of Lazaridis et al. Having said so, it would have been helpful if you had pointed out specific examples where I did that. I think I have been pretty scrupulous about quoting specifically from the text, and rather cautious about extrapolating from it.
As to the validity of the analysis itself, Lazaridis et al is a ground breaking study of the peopling of Europe. I hold it in high esteem, and it seems I am in good company, as 72 of the world's leading population geneticists have signed on as "contributing" authors, including Brenna Henn, Mark Thomas, George Busby, Christian Capelli, Toomas Kivisild, Joachim Burger, Wolfgang Haak (from the group that produced the paper we are discussing) to Qiaomei Fu and Svante Paabo. I hardly think they would have put their names to it if they thought there were serious methodological problems with it. Then, of course, there are the listed co-senior authors, Johannes Kraus and David Reich, although I’m sure as senior authors they have no problems with the conclusions of their own paper. If you wish to pit your expertise in population genetics, the new statistical algorithms that have been developed, and the new statistical tools created to assist in this analysis, that’s of course your prerogative. Far be it from me to suggest that the validity of a study should be determined by an appeal to authority.
The following are just a few thoughts from my non-professional population geneticist’s, “ lay person's” , perspective on the paper and the various tools used in the paper, a lay person who has, however, been reading autosomal analyses and papers since the publication of The History and Geography of Human Genes by Luca Cavalli-Sforza in 1994, (a book that is still invaluable as a primer in this area), so I have, I hope, some familiarity with the issues.
You state that you are concerned that the results from the ADMIXTURE software analysis are not valid because there are not dozens of samples from each population group. More samples are always better. However, autosomal DNA is not like uniparental DNA. There has been so much "mixing" that, from what I have seen, you don't need a lot of samples to get a statistically reliable number for a certain area. There are analyses that address this issue, but I don't have the time to search for them right now. Just as a simple example, there are numerous sample sources for Tuscans. One has 7 samples (HGDP), one has 20 (Hap Map 3) and one has 100 individuals in it (1000 Genomes Project). There is virtually no difference in the autosomal analysis results. For another example, again from the Italian population, the results for the people in the Bergamo HGDP sample are almost identical to the garden variety Northern Italian participants in Dienekes' Dodecad sample.
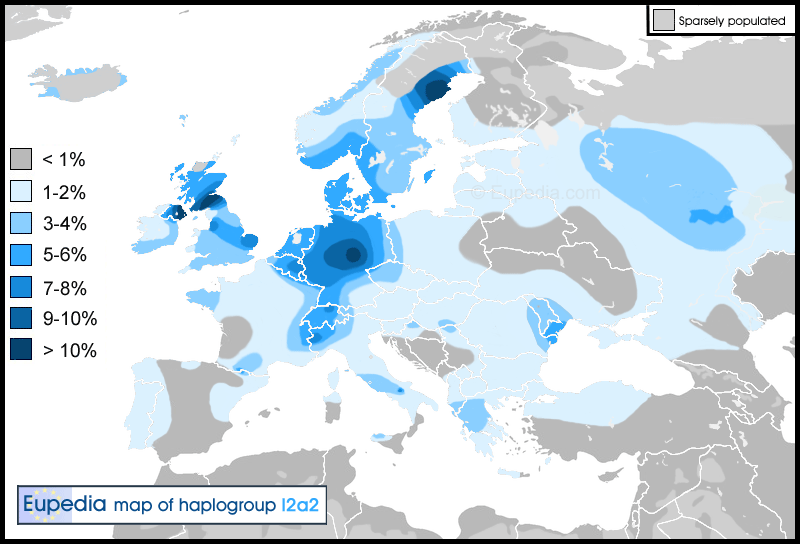
(Also, to your point about uniparental markers, they are good for tracking individual migration movements, but they are very poor indicators, particularly in the case of yDNA, of autosomal make-up, as I would think you would know. There is the often cited example of some pastoralists in western Sub-Saharan Africa who carry very significant amounts of R1b, and yet they are autosomally practically indistinguishable from surrounding non R1b tribes. Particularly as to your point about the ancient genomes, Loschbour is I2a, and La Brana is “C”, a haplogroup we now associate with eastern Asia. Yet, they form an autosomal cluster. )
Further, as to your point about the analysis with ADMIXTURE, if you re-read the paper and the supplement, it might refresh your recollection as to the fact that the Admixture software is just
[FONT="]one [/FONT]of the tools they used to reach their conclusions. They also used PCA analysis, f-statistical analysis and ADMIXTUREGRAPH, unsupervised Mix Mapper and Tree Mix software, plus IBD analysis and chromosome painting analysis to arrive at their conclusions.
.
Now, would it have given the consumers of the paper more information if more populations were tested? The answer is yes, of course. That doesn't invalidate the results, which state broadly that "these analyses allow us to infer that EEF ancestry in Europe today ranges from ~ 30% in the Baltic region to ~90% in the Mediterranean, a gradient which is also consistent with patterns of identity by descent sharing (IBD)(S118) and chromosome painting (S119)”
Those patterns would indicate, in my opinion, that the area from Czechoslovakia (which
[FONT="]was[/FONT] tested-.495 EEF), west through Germany and the Netherlands to Britain(which was also tested and had exactly the same EEF score-.495) hovers around a 49-50% rate on average, a fact that is borne out in the results from the calculator, with the number dropping a few points in far northern Germany from what I recall. (Although I have criticized that calculator, Central Europe, going by the correlation between the calculator results and the study results, is not one of the geographic areas of concern. ) Ukrainians score .46, by the way.
Of course, it’s unfortunate that more western and northwestern European populations were not included. I’m sure that they will be included in the future, since, as the authors speculate, that
may be why their PCA does not exactly duplicate those of some prior studies. However, that is one, and not a particularly good tool, so this doesn’t in any way invalidate the findings of the paper or the percentages of these ancient genomes that show up in modern populations. It just means we don’t have the figures for some modern European populations.
From the figures that
are provided, there seems to be a very narrow range in terms of these scores across the northern European plain and into at least part of the British Isles, which goes from about .46 for Ukrainians for EEF to about .49 in Britain. The number for EEF starts to increase in France, where a more northerly sample is .554 , but France_South is already .675 EEF. The EEF number of about 50% in Central Europe jumps to .715 in northern Italy, .713 in Spanish_North, .712 in Bulgaria and then to .745 in Tuscany, .81 in the Spanish etc. and so on. This all makes total sense in terms of prior autosomal analyses of modern Europeans.
I would be absolutely shocked if there is much difference in the scores for people from different areas on the Northern European plain. That would be contrary to every result I've ever seen from personal or academic autosomal genetic analysis of these groups. There is a great deal of homogeneity in this area, which crosses national borders. Even since the period of the Indo-European migrations, you have the “mixing” that took place in this region as the result of the Germanic migrations, the Slavic migrations, the numerous wars like the Thirty Years War, the migrations brought about by industrialization, and then the dislocations of the First and Second World Wars. Many of these events did
not impact populations in Iberia and the Italian peninsula to the same degree. If you've read all the pertinent papers, and followed the personal results as well, I don't understand how you can fail to see this pattern.
For those who have not seen them, these are some graphics of PCA’s based on autosomal results for Europeans that I think nicely illustrate the point:
http://upload.wikimedia.org/wikiped...tic_structure_(based_on_SNPs)_PC_analysis.png
http://2.bp.blogspot.com/_Ish7688voT0/R6Nt4XlfrwI/AAAAAAAAAFA/NzaQHAnUOvI/s1600-h/pc300k.jpg
http://s21.postimg.org/6pawkhkjb/russiangwafig3.png
You also make some statements that I think are very off the mark, given the reading that I have done on this subject. Why on earth, for example, would it be surprising that Icelanders have an EEF component? They are a mixture of settlers from Norway and Ireland, both areas that received genetic input from Neolithic farmers. (Norway has an EEF score of .41 and Scotland has an EEF score of .39. This makes the Icelandic score of .394 perfectly plausible.) The Neolithic farmers also spread to South Asia (a recent paper explored it in depth) and, yes, Africa, in the form of pastoralism, with some speculation that it's connected to the Cushitic speakers, as per your comment about the Afars of Ethiopia, who are pastoralists who indeed speak a Cushite language. Perhaps you haven't gotten around to reading the papers showing a large Southwest Asian autosomal component in Horner populations, some of which could indeed have entered their genomes with such a migration? Ukrainians also have, of course, an EEF component, as do far NWestern Russians, but the latter so called “ Volga Russian” population is a heavily Siberian admixed population, so their numbers will, of course, be quite different. You may also not be aware of the detail that the Algonquin, Cree and Ojibway are some of the most heavily European admixed populations among all Amerindian groups. There is speculation as well that they have a somewhat differential gene flow as shown by their levels of mtDNA “X”. These are all known relationships genetically speaking.
I also don't quite know how to respond to your complaint that Lazaridis et al doesn't tell us the fate of the WHG in the various areas of Europe. (Although they do, contrary to your assertion, speculate that the perhaps 20% WHG in EEF is attributable to the Mediterranean HGs. They obviously need ancient samples from them to be sure. Also, they did indeed try to fit the SHG into the model, but the statistical analysis refuted it.)
This is an incremental process...as more ancient data becomes available, from Mediterranean HGs, from Ancient Near Eastern farmers, from the first farmers in the Balkans, indeed, from Mesolithic samples
in the Balkans, from Samara and from the new groups entering the Balkans around 3500 BC, I’m sure this group will adjust their analyses, just as they adjusted their findings as published in Moorjani et al in the Patterson et al paper.
The fact remains that, properly understood, this is a quantum leap forward in terms of genetics population studies, and, from what I have seen, is accepted as such in the academic and hobbyist communities.
For those who haven’t been tracking the analyses produced by the Reich Lab, I would suggest reading the following papers in the listed order:
Moorjani et al:
http://www.plosgenetics.org/article/info:doi/10.1371/journal.pgen.1001373
Patterson et al:
http://www.genetics.org/content/early/2012/09/06/genetics.112.145037.full.pdf
Lipson et al:
http://arxiv.org/pdf/1212.2555v2.pdf
The following improvements in statistical analysis appeared in these papers:
- Reich et al, 2009 (Indian Cline; introduction of f-statistics)
- Green et al, 2010 (Neandertal genome; introduction of D-statistics)
- Durand et al 2011 (elaboration on D-statistics)
- Meyer et al, 2012 (high coverage Denisova; enhanced D-statistics)
- Pickrell et al, 2012 (Khoisan origins; use of admixture LD with 1-ref)
- Reich et al. (2012 (Native American origins; multiple waves of admixture)
The following tools were used in the following papers: Patterson et al (ADMIXTOOLS), Loh et al (ALDER), and Moorjani et al (updates to ROLLOFF)
I would also recommend a careful reading of Ralph and Coop:
http://www.plosbiology.org/article/info:doi/10.1371/journal.pbio.1001555
To explore in detail what happened to the hunter-gatherers in each area of Europe, their number, where and when the admixture with farmers took place, whether it was from HG's who remained in place, or from later repopulation from the north, or with the Indo-European migrations, wasn't the
[FONT="]purpose [/FONT]of this paper. On the most basic level, what it is showing is that the genes of the WHGs (Loschbour and La Brana) appear in the genomes of modern Europeans in certain percentages. The same applies to the genes of the Stuttgart sample and the ANE sample. (And the Gok sample and Oetzi himself cluster with Stuttgart) The purpose of the paper is to show that the populations of
[FONT="]Europe[/FONT], not the Middle-East, can best be modelled as a mix of three ancient genome clusters. Other admixtures, when more ancient samples are available, are possible. I can't see how they can be faulted for not answering every detailed question we have about every minor as well as major migration flow and extinction that has taken place in a particular area of Europe over the last thousands of years.
Your comment about them "throwing out" certain groups also shows a fundamental misunderstanding, in my opinion, of this analysis. First of all, the Ukrainians can indeed be fit with the three population model. The Finns, the Mordovians, and the Volga Russians cannot, because they have, in the case of the Finns, for example, according to other autosomal analyses,
[FONT="]at least[/FONT] 5-6% "Eastern" admixture. (Siberian/East Asian, the percentages differing slightly by analysis) They weren't "thrown out". Nor were the Sicilians and the Maltese thrown out. They're all still Europeans. It's just that unlike all other Europeans, they don't fit the model because they have too much "other" admixture, which in the case of the Sicilians and Maltese is not yet properly understood, by the way. Said another way, they are not a statistical fit as a combination of Neolithic farmers from the Near East, Western Hunter Gatherers, and an ancient North Eurasian population that is also ancestral to Amerindians. The fact that the model doesn't fit a few groups at the periphery of Europe doesn't mean that the model is invalid. It
[FONT="]is[/FONT] valid for the vast majority of Europeans.
I don't have the time to address all your points, nor is this the thread for it, but I think I've posted enough to show that I strongly disagree with your analysis. People will have to make up their own minds after reading all the pertinent papers.