The reason for that is because CHG-Iran_Neo and Samara HG do have shared ancestry and if you don't use the supervised mode for your calculator, depending on which population we use first and which we add. The calculator might take this shared ancestry actually as Samara HG instead of CHG or Iran_Neo. There are reasonable arguments from the papers that Samara HG itself could have been already Iran_Neo or CHG admixed. Indeed some samples did show more of this admixture than others.There also seems to be Zarzian and Leyla Tepe culture influence on the Steppe cultures already by Neolithic if not even earlier.
CHG itself is little more ANE and WHG shifted than Iran_Neo, the reason why it shows always some more "EHG" like admixture and in this case East European.
It is archeologically seen also unlikely to assume that a purely Iran_Neo pop crossed into the Steppes without having changed itself slightly on the way up there already. archeoligically the influence makes sense in the way of Iran_CHL => CHL_Caucasus=> Bronze Age Steppes.
This is why the studies gave a ~32% Iran_CHL+~15% CHG like + 53% Samara HG like as a good model, because these Iranian_Plateau herders obviously catched up some additional admxiture on the Caucasus before they mixed with the Samara H&Gs like population, if they even took this route and not the South_Central Asian way were I expect that we will find a Iran_Neo+EHG like people by Neolithic already.
I know what you are saying, they all have ancestral components and might have mixed along the way somewhat too. I believe it is very hard for programmers to really separate mixtures. However, in my observations, I used changes in proportions of admixtures to figure out population movements rather than comparing admixtures alone. Also only dominant admixtures were used to make sure they don't dissipate on their way into a noise.
I do believe Iran-Neo picked up some CHG on their way to the steppe, but it couldn't have been more than 25%. Otherwise we would see proportion of Caucasian to Baloch skewed towards Caucasian.
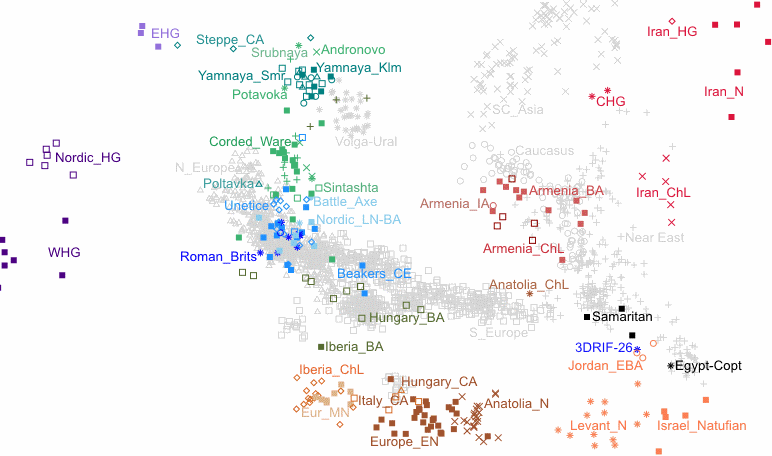
On PCA charts, the distance and direction Yamnaya went from EHG position towards Iranian Neo/CHG, points to 25-30% admixture, not more.
http://eurogenes.blogspot.ca/2016/06/a-moment-of-clarity.html
What you actually saying is very visible in Global 10 K11 ran. They use source population of 100% CHG and 100% Iranian Neolithic. This guys are very related and have probably 90% same genome. In Iranian Neolithic there is 90% of CHG and vice versa, and yet they managed to created 100% source in both. That's quite a paradox.
I'm guessing that they created CHG source population from 100% CHG, and used 10% distinct genome part of Iran Neo to make 100% Iran Neo. Right? I might be wrong.
Anyway, at 10% of total genome it is easy to lose, Iran Neo when it travels to Yamnaya and mixes with locals, getting divided by half and half till it vanishes. Also ancient genomes are not very complete to start with and can easily miss such small parts, which is not helping when dealing with 10% small Iran Neo DNA. In case of K11 Yamnaya is based on almost 50/50 Karelia EHG/Kotias CHG. All hunter gatherers. Anyway, instead of seeing mostly Iranian Neo, which we should to my understanding, we see almost only CHG. Not realising that this CHG is the CHG part in Iranian Neolithic Genome.
Well, maybe I'm wrong, but I think this is what happened here.
Another thing increasing CHG admixture in Yamnaya is that EHG part is based on Karelia and not on Samara. I'm sure Samara being closer had more ancient similarities to CHG than Karelia guy. So a part of Samara EHG is associated with CHG instead of with EHG, making neolithic impact of south caucasus on Yamnaya exaggerated.
Unfortunately I can't confirm it by Gedmatch as the only Karelia sample, which used to be available, can't be found in the system anymore.