I was trying to model some populations and this happened:
Target: Irish
Distance: 4.9374% / 0.04937387
49.8 Yamnaya_RUS_Samara
36.4 Anatolia_Barcin_N
13.8 WHG
Target: Polish olish16
olish16
Distance: 4.6207% / 0.04620698
37.2 Anatolia_Barcin_N
28.6 Yamnaya_RUS_Samara
24.4 RUS_Karelia_HG
9.8 WHG
Target: Estonian
Distance: 6.4167% / 0.06416713
32.0 Anatolia_Barcin_N
30.4 RUS_Karelia_HG
27.2 Yamnaya_RUS_Samara
10.4 WHG
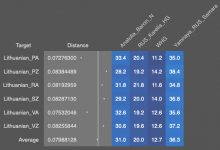
Yamnaya numbers fall in eastern Europe and Karelia HG ( EHG population without Yamnaya CHG mixture) goes up
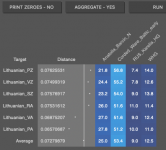
I added 'Corded_Ware_Baltic_early' to the source to see if the model would stop pulling an additional Karelia mix to Eastern Europe. The model has become more efficient and distances have fallen, but eastern Europe continues with an additional mixture of pre-Yamnaya EHG ( it seems to coincide with regions where the R1b haplogroup is currently less common)
Target: Irish
Distance: 4.0625% / 0.04062541
51.8 Corded_Ware_Baltic_early
36.4 Anatolia_Barcin_N
11.8 WHG
Target: Polish: Polish16
Distance: 3.9425% / 0.03942502
37.6 Corded_Ware_Baltic_early
34.8 Anatolia_Barcin_N
17.0 RUS_Karelia_HG
10.6 WHG
Target: Estonian
Distance: 5.7087% / 0.05708720
38.8 Corded_Ware_Baltic_early
28.8 Anatolia_Barcin_N
20.6 RUS_Karelia_HG
11.8 WHG
What could explain the difference?
Target: Irish
Distance: 4.9374% / 0.04937387
49.8 Yamnaya_RUS_Samara
36.4 Anatolia_Barcin_N
13.8 WHG
Target: Polish
Distance: 4.6207% / 0.04620698
37.2 Anatolia_Barcin_N
28.6 Yamnaya_RUS_Samara
24.4 RUS_Karelia_HG
9.8 WHG
Target: Estonian
Distance: 6.4167% / 0.06416713
32.0 Anatolia_Barcin_N
30.4 RUS_Karelia_HG
27.2 Yamnaya_RUS_Samara
10.4 WHG
Yamnaya numbers fall in eastern Europe and Karelia HG ( EHG population without Yamnaya CHG mixture) goes up
I added 'Corded_Ware_Baltic_early' to the source to see if the model would stop pulling an additional Karelia mix to Eastern Europe. The model has become more efficient and distances have fallen, but eastern Europe continues with an additional mixture of pre-Yamnaya EHG ( it seems to coincide with regions where the R1b haplogroup is currently less common)
Target: Irish
Distance: 4.0625% / 0.04062541
51.8 Corded_Ware_Baltic_early
36.4 Anatolia_Barcin_N
11.8 WHG
Target: Polish: Polish16
Distance: 3.9425% / 0.03942502
37.6 Corded_Ware_Baltic_early
34.8 Anatolia_Barcin_N
17.0 RUS_Karelia_HG
10.6 WHG
Target: Estonian
Distance: 5.7087% / 0.05708720
38.8 Corded_Ware_Baltic_early
28.8 Anatolia_Barcin_N
20.6 RUS_Karelia_HG
11.8 WHG
What could explain the difference?