Genomelink’s global ancestry (New): Uses the XGMix algorithm for deeper DNA analysis (at the Chromosome level) that shows how the ancestry breaks down at the chromosome level and also shows the overalls ethnicity estimate. Backed by Carlos D. Bustamante, Alexander Ioanidis, Razib Khan.
see:
https://genomelink.io/product/global-ancestry-dna-report
https://www.biorxiv.org/content/10.1101/2020.04.21.053876v1.full.pdf
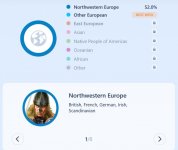
My overall ethnicities



chromosome level (examples of 4 out of 16 ethnicities displayed for me):



(…)

see:
https://genomelink.io/product/global-ancestry-dna-report
https://www.biorxiv.org/content/10.1101/2020.04.21.053876v1.full.pdf
My overall ethnicities



chromosome level (examples of 4 out of 16 ethnicities displayed for me):



(…)

Last edited: