To the admixed haplotypes from the bulk of genomic diversity, we assume that modern humans left Africa somewhere between 80–50 thousand years ago (Kya) and that at that time Neandertals occupied western Eurasia (Stringer 2002; Klein 2003; Mellars 2006; Oppenheimer 2009; Petraglia et al. 2010). Because these populations diverged 400–800 Kya, a number of derived alleles present in the Neandertal DNA can be expected to segregate in all human populations including sub-Saharan Africans. In contrast, segments in human DNA that were subsequently admixed outside Africa are expected to carry additional derived alleles shared with Neandertals but absent in sub-Saharan Africans. The admixed haplotypes are also expected to be younger, that is, less diversified by recombination than the average African haplotypes at the same loci. Ideally, the allelic structure of these haplotypes would differ from the bulk of the common haplotypes reflecting their origin along a separately evolving lineage.
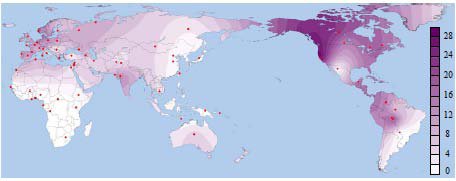
An X-linked haplotype that fulfills these characteristics occurs in an 8-Kb intronic segment spanning exon 44 of the dystrophin gene, referred to as dys44 (Zietkiewicz et al. 1997). We analyzed dys44 polymorphisms in a sample of 6,092 X-chromosomes representing populations from all habitable continents (supplementary table S1, Supplementary Material online), including previously published data (Labuda, Labuda, et al. 2000; Zietkiewicz et al. 2003; Xiao et al. 2004; Lovell et al. 2005; Bourgeois et al. 2009). We focus on the 12 major dys44 haplotypes that explain 89% of genetic diversity in non-Africans and 67% in sub-Saharan Africans (table 1). Of these, haplotype B006 is structurally distinct; with only four derived alleles, it is the closest to the ancestral one. Common outside Africa and virtually absent in sub-Saharan Africa (fig. 1), B006 was earlier proposed to represent an unknown non-African contribution to the human gene pool (Zietkiewicz et al. 2003). Of 1,420 sub-Saharan chromosomes, only one copy of B006 was observed in Ethiopia, and five in Burkina Faso, one among the Rimaibe and four among the Fulani and Tuareg, nomad-pastoralists known for having contacts with northern populations (supplementary table S1, Supplementary Material online). B006 only occurrence at the northern and northeastern outskirts of sub-Saharan Africa is thus likely to be a result of gene flow from a non-African source.
Twelve of the 35 dys44 polymorphisms available in the HapMap3 database are sufficient to identify the 12 major dys44 haplotypes presented in table 1. In the considered HapMap3 populations (supplementary table S1, Supplementary Material online), these haplotypes represent 369 of 523 (70.6%) sub-Saharan chromosomes and 815 of 891 (91.5%) non-African chromosomes that include 77 copies of B006. We extended the dys44 8-kb region by 28 additional polymorphisms on the left and 49 on the right (108 kb in total) to include sites that by inspection appeared to maintain some degree of linkage disequilibrium with the B006 haplotype (fig. 3). In the Neandertal sequence, no information is available for 28 of these sites, 36 sites represent ancestral alleles and 13 derived alleles. Importantly, three of the derived alleles (rs17243319, rs1456729, and rs11796299 in supplementary table S2, Supplementary Material online) are absent from the African chromosomes, as in the case of the derived G of rs11795471 from within B006. Moreover, all derived alleles shared with Neandertals occur at high frequencies (0.75 and more) on a background of the extended B006 haplotype (fig. 3 and supplementary table S2, Supplementary Material online) as expected in a segment of recent Neandertal origin.
https://academic.oup.com/mbe/article/28/7/1957/1048596/An-X-Linked-Haplotype-of-Neandertal-Origin-Is