-
Don't want to see ads? Install an adblocker like uBlock Origin or use a Europe-based privacy-friendly browser like Vivaldi or Mullvad.
You are using an out of date browser. It may not display this or other websites correctly.
You should upgrade or use an alternative browser.
You should upgrade or use an alternative browser.
LivingDNA Living DNA results and comparison
- Thread starter I1a3_Young
- Start date
I1a3_Young
Regular Member
- Messages
- 556
- Reaction score
- 66
- Points
- 28
- Location
- FL
- Ethnic group
- Basically British
- Y-DNA haplogroup
- I1 Z63*
- mtDNA haplogroup
- H5b1
Here's a K36 of repaired vs old Ancestry file:
[TABLE="width: 790"]
[TR]
[TD]Repaired[/TD]
[TD][/TD]
[TD][/TD]
[TD]Old[/TD]
[TD][/TD]
[TD][/TD]
[TD]Delta[/TD]
[/TR]
[TR]
[TD]North_Sea[/TD]
[TD]21.37[/TD]
[TD][/TD]
[TD]North_Sea[/TD]
[TD]21.37[/TD]
[TD][/TD]
[TD="align: right"]0[/TD]
[/TR]
[TR]
[TD]North_Atlantic[/TD]
[TD]13.2[/TD]
[TD][/TD]
[TD]North_Atlantic[/TD]
[TD]13.12[/TD]
[TD][/TD]
[TD="align: right"]0.08[/TD]
[/TR]
[TR]
[TD]Iberian[/TD]
[TD]13[/TD]
[TD][/TD]
[TD]Iberian[/TD]
[TD]12.99[/TD]
[TD][/TD]
[TD="align: right"]0.01[/TD]
[/TR]
[TR]
[TD]Central_Euro[/TD]
[TD]10.59[/TD]
[TD][/TD]
[TD]Central_Euro[/TD]
[TD]10.56[/TD]
[TD][/TD]
[TD="align: right"]0.03[/TD]
[/TR]
[TR]
[TD]Italian[/TD]
[TD]9.01[/TD]
[TD][/TD]
[TD]Italian[/TD]
[TD]9.06[/TD]
[TD][/TD]
[TD="align: right"]-0.05[/TD]
[/TR]
[TR]
[TD]East_Central_Euro[/TD]
[TD]7.66[/TD]
[TD][/TD]
[TD]East_Central_Euro[/TD]
[TD]7.72[/TD]
[TD][/TD]
[TD="align: right"]-0.06[/TD]
[/TR]
[TR]
[TD]Fennoscandian[/TD]
[TD]7.61[/TD]
[TD][/TD]
[TD]Fennoscandian[/TD]
[TD]7.62[/TD]
[TD][/TD]
[TD="align: right"]-0.01[/TD]
[/TR]
[TR]
[TD]French[/TD]
[TD]6.39[/TD]
[TD][/TD]
[TD]French[/TD]
[TD]6.37[/TD]
[TD][/TD]
[TD="align: right"]0.02[/TD]
[/TR]
[TR]
[TD]Basque[/TD]
[TD]4.38[/TD]
[TD][/TD]
[TD]Basque[/TD]
[TD]4.39[/TD]
[TD][/TD]
[TD="align: right"]-0.01[/TD]
[/TR]
[TR]
[TD]Volga-Ural[/TD]
[TD]2.59[/TD]
[TD][/TD]
[TD]Volga-Ural[/TD]
[TD]2.56[/TD]
[TD][/TD]
[TD="align: right"]0.03[/TD]
[/TR]
[TR]
[TD]West_Caucasian[/TD]
[TD]2.22[/TD]
[TD][/TD]
[TD]West_Caucasian[/TD]
[TD]2.24[/TD]
[TD][/TD]
[TD="align: right"]-0.02[/TD]
[/TR]
[TR]
[TD]North_Caucasian[/TD]
[TD]1.58[/TD]
[TD][/TD]
[TD]North_Caucasian[/TD]
[TD]1.57[/TD]
[TD][/TD]
[TD="align: right"]0.01[/TD]
[/TR]
[TR]
[TD]West_Med[/TD]
[TD]0.39[/TD]
[TD][/TD]
[TD]West_Med[/TD]
[TD]0.4[/TD]
[TD][/TD]
[TD="align: right"]-0.01[/TD]
[/TR]
[/TABLE]
No, Davef I have not had a 23andMe test. I have only had Ancestry + LivingDNA tests plus free uploads like MyHeritage and GEDmatch.
[TABLE="width: 790"]
[TR]
[TD]Repaired[/TD]
[TD][/TD]
[TD][/TD]
[TD]Old[/TD]
[TD][/TD]
[TD][/TD]
[TD]Delta[/TD]
[/TR]
[TR]
[TD]North_Sea[/TD]
[TD]21.37[/TD]
[TD][/TD]
[TD]North_Sea[/TD]
[TD]21.37[/TD]
[TD][/TD]
[TD="align: right"]0[/TD]
[/TR]
[TR]
[TD]North_Atlantic[/TD]
[TD]13.2[/TD]
[TD][/TD]
[TD]North_Atlantic[/TD]
[TD]13.12[/TD]
[TD][/TD]
[TD="align: right"]0.08[/TD]
[/TR]
[TR]
[TD]Iberian[/TD]
[TD]13[/TD]
[TD][/TD]
[TD]Iberian[/TD]
[TD]12.99[/TD]
[TD][/TD]
[TD="align: right"]0.01[/TD]
[/TR]
[TR]
[TD]Central_Euro[/TD]
[TD]10.59[/TD]
[TD][/TD]
[TD]Central_Euro[/TD]
[TD]10.56[/TD]
[TD][/TD]
[TD="align: right"]0.03[/TD]
[/TR]
[TR]
[TD]Italian[/TD]
[TD]9.01[/TD]
[TD][/TD]
[TD]Italian[/TD]
[TD]9.06[/TD]
[TD][/TD]
[TD="align: right"]-0.05[/TD]
[/TR]
[TR]
[TD]East_Central_Euro[/TD]
[TD]7.66[/TD]
[TD][/TD]
[TD]East_Central_Euro[/TD]
[TD]7.72[/TD]
[TD][/TD]
[TD="align: right"]-0.06[/TD]
[/TR]
[TR]
[TD]Fennoscandian[/TD]
[TD]7.61[/TD]
[TD][/TD]
[TD]Fennoscandian[/TD]
[TD]7.62[/TD]
[TD][/TD]
[TD="align: right"]-0.01[/TD]
[/TR]
[TR]
[TD]French[/TD]
[TD]6.39[/TD]
[TD][/TD]
[TD]French[/TD]
[TD]6.37[/TD]
[TD][/TD]
[TD="align: right"]0.02[/TD]
[/TR]
[TR]
[TD]Basque[/TD]
[TD]4.38[/TD]
[TD][/TD]
[TD]Basque[/TD]
[TD]4.39[/TD]
[TD][/TD]
[TD="align: right"]-0.01[/TD]
[/TR]
[TR]
[TD]Volga-Ural[/TD]
[TD]2.59[/TD]
[TD][/TD]
[TD]Volga-Ural[/TD]
[TD]2.56[/TD]
[TD][/TD]
[TD="align: right"]0.03[/TD]
[/TR]
[TR]
[TD]West_Caucasian[/TD]
[TD]2.22[/TD]
[TD][/TD]
[TD]West_Caucasian[/TD]
[TD]2.24[/TD]
[TD][/TD]
[TD="align: right"]-0.02[/TD]
[/TR]
[TR]
[TD]North_Caucasian[/TD]
[TD]1.58[/TD]
[TD][/TD]
[TD]North_Caucasian[/TD]
[TD]1.57[/TD]
[TD][/TD]
[TD="align: right"]0.01[/TD]
[/TR]
[TR]
[TD]West_Med[/TD]
[TD]0.39[/TD]
[TD][/TD]
[TD]West_Med[/TD]
[TD]0.4[/TD]
[TD][/TD]
[TD="align: right"]-0.01[/TD]
[/TR]
[/TABLE]
No, Davef I have not had a 23andMe test. I have only had Ancestry + LivingDNA tests plus free uploads like MyHeritage and GEDmatch.
Johane Derite
Regular Member
- Messages
- 1,959
- Reaction score
- 1,057
- Points
- 113
- Ethnic group
- Albanian
- Y-DNA haplogroup
- E-V13>Z5018>FGC33625
- mtDNA haplogroup
- U1a1a
Here's a K36 of repaired vs old Ancestry file:
Pretty close. 1st question, how did you repair the livingdna raw data.
2nd question, have you had any correspondence with livingdna over their issues with the raw data? Have they indicated at all
whether they will update their raw file formats or does it seem that this is their final offer?
thx
I1a3_Young
Regular Member
- Messages
- 556
- Reaction score
- 66
- Points
- 28
- Location
- FL
- Ethnic group
- Basically British
- Y-DNA haplogroup
- I1 Z63*
- mtDNA haplogroup
- H5b1
Pretty close. 1st question, how did you repair the livingdna raw data.
2nd question, have you had any correspondence with livingdna over their issues with the raw data? Have they indicated at all
whether they will update their raw file formats or does it seem that this is their final offer?
thx
I used Microsoft Excel to split genotype columns to match Ancestry raw format. I sorted for bad values and then used =VLOOKUP() to find the corresponding values for the bad positions (both ways). Then for the LivingDNA I recombined the columns to put both alleles back together and pasted over the bad values.
I haven't been able to successfully upload the repaired LDNA file yet, I get an error. The repaired Ancestry did pull my numbers into more correct territory ever so slightly.
Edit: I saved the file in Apple format .txt instead of tab delimited or MSDOS and it appears to be working. I'll have repaired LivingDNA figures to compare to repaired Ancestry.com figures!
I1a3_Young
Regular Member
- Messages
- 556
- Reaction score
- 66
- Points
- 28
- Location
- FL
- Ethnic group
- Basically British
- Y-DNA haplogroup
- I1 Z63*
- mtDNA haplogroup
- H5b1
[TABLE="width: 653"]
[TR]
[TD]#[/TD]
[TD]Population[/TD]
[TD]Percent[/TD]
[TD][/TD]
[TD]#[/TD]
[TD]Population[/TD]
[TD]Percent[/TD]
[/TR]
[TR]
[TD]1[/TD]
[TD]North_Atlantic[/TD]
[TD]49.39[/TD]
[TD][/TD]
[TD]1[/TD]
[TD]North_Atlantic[/TD]
[TD]49.76[/TD]
[/TR]
[TR]
[TD]2[/TD]
[TD]Baltic[/TD]
[TD]23.47[/TD]
[TD][/TD]
[TD]2[/TD]
[TD]Baltic[/TD]
[TD]23.09[/TD]
[/TR]
[TR]
[TD]3[/TD]
[TD]West_Med[/TD]
[TD]12.48[/TD]
[TD][/TD]
[TD]3[/TD]
[TD]West_Med[/TD]
[TD]12.56[/TD]
[/TR]
[TR]
[TD]4[/TD]
[TD]East_Med[/TD]
[TD]5.11[/TD]
[TD][/TD]
[TD]4[/TD]
[TD]East_Med[/TD]
[TD]5.25[/TD]
[/TR]
[TR]
[TD]5[/TD]
[TD]West_Asian[/TD]
[TD]5.11[/TD]
[TD][/TD]
[TD]5[/TD]
[TD]West_Asian[/TD]
[TD]5.07[/TD]
[/TR]
[TR]
[TD]6[/TD]
[TD]Red_Sea[/TD]
[TD]1.54[/TD]
[TD][/TD]
[TD]6[/TD]
[TD]Red_Sea[/TD]
[TD]1.46[/TD]
[/TR]
[TR]
[TD]7[/TD]
[TD]South_Asian[/TD]
[TD]1.16[/TD]
[TD][/TD]
[TD]7[/TD]
[TD]South_Asian[/TD]
[TD]1.29[/TD]
[/TR]
[TR]
[TD]8[/TD]
[TD]Northeast_African[/TD]
[TD]0.64[/TD]
[TD][/TD]
[TD]8[/TD]
[TD]Northeast_African[/TD]
[TD]0.59[/TD]
[/TR]
[TR]
[TD]9[/TD]
[TD]Sub-Saharan[/TD]
[TD]0.61[/TD]
[TD][/TD]
[TD]9[/TD]
[TD]Oceanian[/TD]
[TD]0.42[/TD]
[/TR]
[TR]
[TD]10[/TD]
[TD]Oceanian[/TD]
[TD]0.5[/TD]
[TD][/TD]
[TD]10[/TD]
[TD]Sub-Saharan[/TD]
[TD]0.37[/TD]
[/TR]
[TR]
[TD][/TD]
[TD][/TD]
[TD][/TD]
[TD][/TD]
[TD]11[/TD]
[TD]Amerindian[/TD]
[TD]0.13[/TD]
[/TR]
[TR]
[TD]#[/TD]
[TD]Population (source)[/TD]
[TD]Distance[/TD]
[TD][/TD]
[TD]#[/TD]
[TD]Population (source)[/TD]
[TD]Distance[/TD]
[/TR]
[TR]
[TD]1[/TD]
[TD]Southeast_English[/TD]
[TD]2.5[/TD]
[TD][/TD]
[TD]1[/TD]
[TD]Southeast_English[/TD]
[TD]2.2[/TD]
[/TR]
[TR]
[TD]2[/TD]
[TD]Orcadian[/TD]
[TD]3.55[/TD]
[TD][/TD]
[TD]2[/TD]
[TD]Orcadian[/TD]
[TD]3.5[/TD]
[/TR]
[TR]
[TD]3[/TD]
[TD]North_Dutch[/TD]
[TD]3.78[/TD]
[TD][/TD]
[TD]3[/TD]
[TD]Southwest_English[/TD]
[TD]3.88[/TD]
[/TR]
[TR]
[TD]4[/TD]
[TD]Danish[/TD]
[TD]4.01[/TD]
[TD][/TD]
[TD]4[/TD]
[TD]North_Dutch[/TD]
[TD]3.99[/TD]
[/TR]
[TR]
[TD]5[/TD]
[TD]Southwest_English[/TD]
[TD]4.15[/TD]
[TD][/TD]
[TD]5[/TD]
[TD]Danish[/TD]
[TD]4.21[/TD]
[/TR]
[TR]
[TD]6[/TD]
[TD]Irish[/TD]
[TD]4.44[/TD]
[TD][/TD]
[TD]6[/TD]
[TD]Irish[/TD]
[TD]4.34[/TD]
[/TR]
[TR]
[TD]7[/TD]
[TD]West_Scottish[/TD]
[TD]4.64[/TD]
[TD][/TD]
[TD]7[/TD]
[TD]West_Scottish[/TD]
[TD]4.44[/TD]
[/TR]
[TR]
[TD]8[/TD]
[TD]North_German[/TD]
[TD]4.91[/TD]
[TD][/TD]
[TD]8[/TD]
[TD]North_German[/TD]
[TD]5.29[/TD]
[/TR]
[TR]
[TD]9[/TD]
[TD]South_Dutch[/TD]
[TD]5.78[/TD]
[TD][/TD]
[TD]9[/TD]
[TD]South_Dutch[/TD]
[TD]5.85[/TD]
[/TR]
[TR]
[TD]10[/TD]
[TD]Norwegian[/TD]
[TD]6.41[/TD]
[TD][/TD]
[TD]10[/TD]
[TD]Norwegian[/TD]
[TD]6.62[/TD]
[/TR]
[/TABLE]
These are the unrepaired LDNA numbers for EuroK13 from regular GEDMatch (first set) and GENESIS BETA (second set). Same exact file.
[TR]
[TD]#[/TD]
[TD]Population[/TD]
[TD]Percent[/TD]
[TD][/TD]
[TD]#[/TD]
[TD]Population[/TD]
[TD]Percent[/TD]
[/TR]
[TR]
[TD]1[/TD]
[TD]North_Atlantic[/TD]
[TD]49.39[/TD]
[TD][/TD]
[TD]1[/TD]
[TD]North_Atlantic[/TD]
[TD]49.76[/TD]
[/TR]
[TR]
[TD]2[/TD]
[TD]Baltic[/TD]
[TD]23.47[/TD]
[TD][/TD]
[TD]2[/TD]
[TD]Baltic[/TD]
[TD]23.09[/TD]
[/TR]
[TR]
[TD]3[/TD]
[TD]West_Med[/TD]
[TD]12.48[/TD]
[TD][/TD]
[TD]3[/TD]
[TD]West_Med[/TD]
[TD]12.56[/TD]
[/TR]
[TR]
[TD]4[/TD]
[TD]East_Med[/TD]
[TD]5.11[/TD]
[TD][/TD]
[TD]4[/TD]
[TD]East_Med[/TD]
[TD]5.25[/TD]
[/TR]
[TR]
[TD]5[/TD]
[TD]West_Asian[/TD]
[TD]5.11[/TD]
[TD][/TD]
[TD]5[/TD]
[TD]West_Asian[/TD]
[TD]5.07[/TD]
[/TR]
[TR]
[TD]6[/TD]
[TD]Red_Sea[/TD]
[TD]1.54[/TD]
[TD][/TD]
[TD]6[/TD]
[TD]Red_Sea[/TD]
[TD]1.46[/TD]
[/TR]
[TR]
[TD]7[/TD]
[TD]South_Asian[/TD]
[TD]1.16[/TD]
[TD][/TD]
[TD]7[/TD]
[TD]South_Asian[/TD]
[TD]1.29[/TD]
[/TR]
[TR]
[TD]8[/TD]
[TD]Northeast_African[/TD]
[TD]0.64[/TD]
[TD][/TD]
[TD]8[/TD]
[TD]Northeast_African[/TD]
[TD]0.59[/TD]
[/TR]
[TR]
[TD]9[/TD]
[TD]Sub-Saharan[/TD]
[TD]0.61[/TD]
[TD][/TD]
[TD]9[/TD]
[TD]Oceanian[/TD]
[TD]0.42[/TD]
[/TR]
[TR]
[TD]10[/TD]
[TD]Oceanian[/TD]
[TD]0.5[/TD]
[TD][/TD]
[TD]10[/TD]
[TD]Sub-Saharan[/TD]
[TD]0.37[/TD]
[/TR]
[TR]
[TD][/TD]
[TD][/TD]
[TD][/TD]
[TD][/TD]
[TD]11[/TD]
[TD]Amerindian[/TD]
[TD]0.13[/TD]
[/TR]
[TR]
[TD]#[/TD]
[TD]Population (source)[/TD]
[TD]Distance[/TD]
[TD][/TD]
[TD]#[/TD]
[TD]Population (source)[/TD]
[TD]Distance[/TD]
[/TR]
[TR]
[TD]1[/TD]
[TD]Southeast_English[/TD]
[TD]2.5[/TD]
[TD][/TD]
[TD]1[/TD]
[TD]Southeast_English[/TD]
[TD]2.2[/TD]
[/TR]
[TR]
[TD]2[/TD]
[TD]Orcadian[/TD]
[TD]3.55[/TD]
[TD][/TD]
[TD]2[/TD]
[TD]Orcadian[/TD]
[TD]3.5[/TD]
[/TR]
[TR]
[TD]3[/TD]
[TD]North_Dutch[/TD]
[TD]3.78[/TD]
[TD][/TD]
[TD]3[/TD]
[TD]Southwest_English[/TD]
[TD]3.88[/TD]
[/TR]
[TR]
[TD]4[/TD]
[TD]Danish[/TD]
[TD]4.01[/TD]
[TD][/TD]
[TD]4[/TD]
[TD]North_Dutch[/TD]
[TD]3.99[/TD]
[/TR]
[TR]
[TD]5[/TD]
[TD]Southwest_English[/TD]
[TD]4.15[/TD]
[TD][/TD]
[TD]5[/TD]
[TD]Danish[/TD]
[TD]4.21[/TD]
[/TR]
[TR]
[TD]6[/TD]
[TD]Irish[/TD]
[TD]4.44[/TD]
[TD][/TD]
[TD]6[/TD]
[TD]Irish[/TD]
[TD]4.34[/TD]
[/TR]
[TR]
[TD]7[/TD]
[TD]West_Scottish[/TD]
[TD]4.64[/TD]
[TD][/TD]
[TD]7[/TD]
[TD]West_Scottish[/TD]
[TD]4.44[/TD]
[/TR]
[TR]
[TD]8[/TD]
[TD]North_German[/TD]
[TD]4.91[/TD]
[TD][/TD]
[TD]8[/TD]
[TD]North_German[/TD]
[TD]5.29[/TD]
[/TR]
[TR]
[TD]9[/TD]
[TD]South_Dutch[/TD]
[TD]5.78[/TD]
[TD][/TD]
[TD]9[/TD]
[TD]South_Dutch[/TD]
[TD]5.85[/TD]
[/TR]
[TR]
[TD]10[/TD]
[TD]Norwegian[/TD]
[TD]6.41[/TD]
[TD][/TD]
[TD]10[/TD]
[TD]Norwegian[/TD]
[TD]6.62[/TD]
[/TR]
[/TABLE]
These are the unrepaired LDNA numbers for EuroK13 from regular GEDMatch (first set) and GENESIS BETA (second set). Same exact file.
I1a3_Young
Regular Member
- Messages
- 556
- Reaction score
- 66
- Points
- 28
- Location
- FL
- Ethnic group
- Basically British
- Y-DNA haplogroup
- I1 Z63*
- mtDNA haplogroup
- H5b1
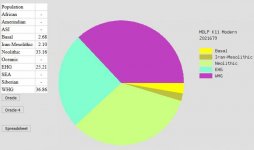
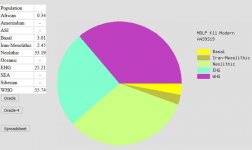
Alright, I have repaired files for Ancestry and LivingDNA uploaded to GENESIS BETA:
This is EurogenesK36. Repairing the LDNA file removed a 0.37 Central African component
[TABLE="width: 290"]
[TR]
[TD]EuroK36 Genesis[/TD]
[TD]Ancest.[/TD]
[TD]LDNA[/TD]
[TD]Delta[/TD]
[/TR]
[TR]
[TD]North_Sea[/TD]
[TD]21.37[/TD]
[TD]23.63[/TD]
[TD="align: right"]2.26[/TD]
[/TR]
[TR]
[TD]North_Atlantic[/TD]
[TD]13.2[/TD]
[TD]8.89[/TD]
[TD="align: right"]-4.31[/TD]
[/TR]
[TR]
[TD]Iberian[/TD]
[TD]13[/TD]
[TD]11.67[/TD]
[TD="align: right"]-1.33[/TD]
[/TR]
[TR]
[TD]Central_Euro[/TD]
[TD]10.59[/TD]
[TD]14.83[/TD]
[TD="align: right"]4.24[/TD]
[/TR]
[TR]
[TD]Italian[/TD]
[TD]9.01[/TD]
[TD]6.51[/TD]
[TD="align: right"]-2.5[/TD]
[/TR]
[TR]
[TD]East_Central_Euro[/TD]
[TD]7.66[/TD]
[TD]6.04[/TD]
[TD="align: right"]-1.62[/TD]
[/TR]
[TR]
[TD]Fennoscandian[/TD]
[TD]7.61[/TD]
[TD]6.99[/TD]
[TD="align: right"]-0.62[/TD]
[/TR]
[TR]
[TD]French[/TD]
[TD]6.39[/TD]
[TD]7.77[/TD]
[TD="align: right"]1.38[/TD]
[/TR]
[TR]
[TD]Basque[/TD]
[TD]4.38[/TD]
[TD]5.6[/TD]
[TD="align: right"]1.22[/TD]
[/TR]
[TR]
[TD]Volga-Ural[/TD]
[TD]2.59[/TD]
[TD]1.74[/TD]
[TD="align: right"]-0.85[/TD]
[/TR]
[TR]
[TD]West_Caucasian[/TD]
[TD]2.22[/TD]
[TD]0.56[/TD]
[TD="align: right"]-1.66[/TD]
[/TR]
[TR]
[TD]North_Caucasian[/TD]
[TD]1.58[/TD]
[TD]0[/TD]
[TD="align: right"]-1.58[/TD]
[/TR]
[TR]
[TD]West_Med[/TD]
[TD]0.39[/TD]
[TD]0[/TD]
[TD="align: right"]-0.39[/TD]
[/TR]
[TR]
[TD]Near_Eastern[/TD]
[TD]0[/TD]
[TD]3.98[/TD]
[TD="align: right"]3.98[/TD]
[/TR]
[TR]
[TD]Eastern_Euro[/TD]
[TD]0[/TD]
[TD]1.29[/TD]
[TD="align: right"]1.29[/TD]
[/TR]
[TR]
[TD]North_African[/TD]
[TD]0[/TD]
[TD]0.47[/TD]
[TD="align: right"]0.47[/TD]
[/TR]
[/TABLE]
This is EurogenesK36. Repairing the LDNA file removed a 0.37 Central African component
[TABLE="width: 290"]
[TR]
[TD]EuroK36 Genesis[/TD]
[TD]Ancest.[/TD]
[TD]LDNA[/TD]
[TD]Delta[/TD]
[/TR]
[TR]
[TD]North_Sea[/TD]
[TD]21.37[/TD]
[TD]23.63[/TD]
[TD="align: right"]2.26[/TD]
[/TR]
[TR]
[TD]North_Atlantic[/TD]
[TD]13.2[/TD]
[TD]8.89[/TD]
[TD="align: right"]-4.31[/TD]
[/TR]
[TR]
[TD]Iberian[/TD]
[TD]13[/TD]
[TD]11.67[/TD]
[TD="align: right"]-1.33[/TD]
[/TR]
[TR]
[TD]Central_Euro[/TD]
[TD]10.59[/TD]
[TD]14.83[/TD]
[TD="align: right"]4.24[/TD]
[/TR]
[TR]
[TD]Italian[/TD]
[TD]9.01[/TD]
[TD]6.51[/TD]
[TD="align: right"]-2.5[/TD]
[/TR]
[TR]
[TD]East_Central_Euro[/TD]
[TD]7.66[/TD]
[TD]6.04[/TD]
[TD="align: right"]-1.62[/TD]
[/TR]
[TR]
[TD]Fennoscandian[/TD]
[TD]7.61[/TD]
[TD]6.99[/TD]
[TD="align: right"]-0.62[/TD]
[/TR]
[TR]
[TD]French[/TD]
[TD]6.39[/TD]
[TD]7.77[/TD]
[TD="align: right"]1.38[/TD]
[/TR]
[TR]
[TD]Basque[/TD]
[TD]4.38[/TD]
[TD]5.6[/TD]
[TD="align: right"]1.22[/TD]
[/TR]
[TR]
[TD]Volga-Ural[/TD]
[TD]2.59[/TD]
[TD]1.74[/TD]
[TD="align: right"]-0.85[/TD]
[/TR]
[TR]
[TD]West_Caucasian[/TD]
[TD]2.22[/TD]
[TD]0.56[/TD]
[TD="align: right"]-1.66[/TD]
[/TR]
[TR]
[TD]North_Caucasian[/TD]
[TD]1.58[/TD]
[TD]0[/TD]
[TD="align: right"]-1.58[/TD]
[/TR]
[TR]
[TD]West_Med[/TD]
[TD]0.39[/TD]
[TD]0[/TD]
[TD="align: right"]-0.39[/TD]
[/TR]
[TR]
[TD]Near_Eastern[/TD]
[TD]0[/TD]
[TD]3.98[/TD]
[TD="align: right"]3.98[/TD]
[/TR]
[TR]
[TD]Eastern_Euro[/TD]
[TD]0[/TD]
[TD]1.29[/TD]
[TD="align: right"]1.29[/TD]
[/TR]
[TR]
[TD]North_African[/TD]
[TD]0[/TD]
[TD]0.47[/TD]
[TD="align: right"]0.47[/TD]
[/TR]
[/TABLE]
I1a3_Young
Regular Member
- Messages
- 556
- Reaction score
- 66
- Points
- 28
- Location
- FL
- Ethnic group
- Basically British
- Y-DNA haplogroup
- I1 Z63*
- mtDNA haplogroup
- H5b1
Here is repaired file of Ancestry and LivingDNA using Genesis Beta K13. Notice population match for LDNA is all the way down to 2.2 for SE English. Big difference.
[TABLE="width: 337"]
[TR]
[TD="colspan: 2"]K13 Genesis Beta[/TD]
[TD]AncREP[/TD]
[TD]LDNA REP[/TD]
[/TR]
[TR]
[TD]#[/TD]
[TD]Population[/TD]
[TD]Percent[/TD]
[TD][/TD]
[/TR]
[TR]
[TD]1[/TD]
[TD]North_Atlantic[/TD]
[TD]48.96[/TD]
[TD]49.76[/TD]
[/TR]
[TR]
[TD]2[/TD]
[TD]Baltic[/TD]
[TD]22.24[/TD]
[TD]23.09[/TD]
[/TR]
[TR]
[TD]3[/TD]
[TD]West_Med[/TD]
[TD]13.09[/TD]
[TD]12.56[/TD]
[/TR]
[TR]
[TD]4[/TD]
[TD]West_Asian[/TD]
[TD]7.72[/TD]
[TD]5.07[/TD]
[/TR]
[TR]
[TD]5[/TD]
[TD]East_Med[/TD]
[TD]3.62[/TD]
[TD]5.25[/TD]
[/TR]
[TR]
[TD]6[/TD]
[TD]Red_Sea[/TD]
[TD]2.19[/TD]
[TD]1.46[/TD]
[/TR]
[TR]
[TD]7[/TD]
[TD]Oceanian[/TD]
[TD]0.78[/TD]
[TD]0.42[/TD]
[/TR]
[TR]
[TD]8[/TD]
[TD]Siberian[/TD]
[TD]0.5[/TD]
[TD]0[/TD]
[/TR]
[TR]
[TD]9[/TD]
[TD]Amerindian[/TD]
[TD]0.46[/TD]
[TD]0.13[/TD]
[/TR]
[TR]
[TD]10[/TD]
[TD]Sub-Saharan[/TD]
[TD]0.44[/TD]
[TD]0.37[/TD]
[/TR]
[TR]
[TD]11[/TD]
[TD]South_Asian[/TD]
[TD]0[/TD]
[TD]1.29[/TD]
[/TR]
[TR]
[TD]12[/TD]
[TD]Northeast_African[/TD]
[TD]0[/TD]
[TD]0.59[/TD]
[/TR]
[TR]
[TD]#[/TD]
[TD]Population (source)[/TD]
[TD]Distance[/TD]
[TD][/TD]
[/TR]
[TR]
[TD]1[/TD]
[TD]Southeast_English[/TD]
[TD]4.25[/TD]
[TD]2.2[/TD]
[/TR]
[TR]
[TD]2[/TD]
[TD]Southwest_English[/TD]
[TD]4.38[/TD]
[TD]3.88[/TD]
[/TR]
[TR]
[TD]3[/TD]
[TD]Orcadian[/TD]
[TD]4.42[/TD]
[TD]3.5[/TD]
[/TR]
[TR]
[TD]4[/TD]
[TD]Irish[/TD]
[TD]4.44[/TD]
[TD]4.34[/TD]
[/TR]
[TR]
[TD]5[/TD]
[TD]North_Dutch[/TD]
[TD]4.87[/TD]
[TD]3.99[/TD]
[/TR]
[TR]
[TD]6[/TD]
[TD]West_Scottish[/TD]
[TD]4.91[/TD]
[TD]4.44[/TD]
[/TR]
[TR]
[TD]7[/TD]
[TD]Danish[/TD]
[TD]5.63[/TD]
[TD]4.21[/TD]
[/TR]
[TR]
[TD]8[/TD]
[TD]North_German[/TD]
[TD]5.81[/TD]
[TD]5.29[/TD]
[/TR]
[TR]
[TD]9[/TD]
[TD]South_Dutch[/TD]
[TD]5.92[/TD]
[TD]5.85[/TD]
[/TR]
[TR]
[TD]10[/TD]
[TD]West_German[/TD]
[TD]6.97[/TD]
[TD][/TD]
[/TR]
[TR]
[TD]11[/TD]
[TD]Norwegian[/TD]
[TD]7.33[/TD]
[TD]6.62[/TD]
[/TR]
[TR]
[TD]12[/TD]
[TD]Swedish[/TD]
[TD]9.24[/TD]
[TD][/TD]
[/TR]
[/TABLE]
[TABLE="width: 337"]
[TR]
[TD="colspan: 2"]K13 Genesis Beta[/TD]
[TD]AncREP[/TD]
[TD]LDNA REP[/TD]
[/TR]
[TR]
[TD]#[/TD]
[TD]Population[/TD]
[TD]Percent[/TD]
[TD][/TD]
[/TR]
[TR]
[TD]1[/TD]
[TD]North_Atlantic[/TD]
[TD]48.96[/TD]
[TD]49.76[/TD]
[/TR]
[TR]
[TD]2[/TD]
[TD]Baltic[/TD]
[TD]22.24[/TD]
[TD]23.09[/TD]
[/TR]
[TR]
[TD]3[/TD]
[TD]West_Med[/TD]
[TD]13.09[/TD]
[TD]12.56[/TD]
[/TR]
[TR]
[TD]4[/TD]
[TD]West_Asian[/TD]
[TD]7.72[/TD]
[TD]5.07[/TD]
[/TR]
[TR]
[TD]5[/TD]
[TD]East_Med[/TD]
[TD]3.62[/TD]
[TD]5.25[/TD]
[/TR]
[TR]
[TD]6[/TD]
[TD]Red_Sea[/TD]
[TD]2.19[/TD]
[TD]1.46[/TD]
[/TR]
[TR]
[TD]7[/TD]
[TD]Oceanian[/TD]
[TD]0.78[/TD]
[TD]0.42[/TD]
[/TR]
[TR]
[TD]8[/TD]
[TD]Siberian[/TD]
[TD]0.5[/TD]
[TD]0[/TD]
[/TR]
[TR]
[TD]9[/TD]
[TD]Amerindian[/TD]
[TD]0.46[/TD]
[TD]0.13[/TD]
[/TR]
[TR]
[TD]10[/TD]
[TD]Sub-Saharan[/TD]
[TD]0.44[/TD]
[TD]0.37[/TD]
[/TR]
[TR]
[TD]11[/TD]
[TD]South_Asian[/TD]
[TD]0[/TD]
[TD]1.29[/TD]
[/TR]
[TR]
[TD]12[/TD]
[TD]Northeast_African[/TD]
[TD]0[/TD]
[TD]0.59[/TD]
[/TR]
[TR]
[TD]#[/TD]
[TD]Population (source)[/TD]
[TD]Distance[/TD]
[TD][/TD]
[/TR]
[TR]
[TD]1[/TD]
[TD]Southeast_English[/TD]
[TD]4.25[/TD]
[TD]2.2[/TD]
[/TR]
[TR]
[TD]2[/TD]
[TD]Southwest_English[/TD]
[TD]4.38[/TD]
[TD]3.88[/TD]
[/TR]
[TR]
[TD]3[/TD]
[TD]Orcadian[/TD]
[TD]4.42[/TD]
[TD]3.5[/TD]
[/TR]
[TR]
[TD]4[/TD]
[TD]Irish[/TD]
[TD]4.44[/TD]
[TD]4.34[/TD]
[/TR]
[TR]
[TD]5[/TD]
[TD]North_Dutch[/TD]
[TD]4.87[/TD]
[TD]3.99[/TD]
[/TR]
[TR]
[TD]6[/TD]
[TD]West_Scottish[/TD]
[TD]4.91[/TD]
[TD]4.44[/TD]
[/TR]
[TR]
[TD]7[/TD]
[TD]Danish[/TD]
[TD]5.63[/TD]
[TD]4.21[/TD]
[/TR]
[TR]
[TD]8[/TD]
[TD]North_German[/TD]
[TD]5.81[/TD]
[TD]5.29[/TD]
[/TR]
[TR]
[TD]9[/TD]
[TD]South_Dutch[/TD]
[TD]5.92[/TD]
[TD]5.85[/TD]
[/TR]
[TR]
[TD]10[/TD]
[TD]West_German[/TD]
[TD]6.97[/TD]
[TD][/TD]
[/TR]
[TR]
[TD]11[/TD]
[TD]Norwegian[/TD]
[TD]7.33[/TD]
[TD]6.62[/TD]
[/TR]
[TR]
[TD]12[/TD]
[TD]Swedish[/TD]
[TD]9.24[/TD]
[TD][/TD]
[/TR]
[/TABLE]
Johane Derite
Regular Member
- Messages
- 1,959
- Reaction score
- 1,057
- Points
- 113
- Ethnic group
- Albanian
- Y-DNA haplogroup
- E-V13>Z5018>FGC33625
- mtDNA haplogroup
- U1a1a
Here is repaired file of Ancestry and LivingDNA using Genesis Beta K13. Notice population match for LDNA is all the way down to 2.2 for SE English. Big difference
Damn, it looks i'm not gonna be buying a livingdna to be honest. Have they been responsive at all to your results and issues with the rawdna?
I1a3_Young
Regular Member
- Messages
- 556
- Reaction score
- 66
- Points
- 28
- Location
- FL
- Ethnic group
- Basically British
- Y-DNA haplogroup
- I1 Z63*
- mtDNA haplogroup
- H5b1
Damn, it looks i'm not gonna be buying a livingdna to be honest. Have they been responsive at all to your results and issues with the rawdna?
I haven't asked them anything. My questions will be about the "Kurdish" and whether I can get the Y positions tested in addition to the positive Y calls.
Johane Derite
Regular Member
- Messages
- 1,959
- Reaction score
- 1,057
- Points
- 113
- Ethnic group
- Albanian
- Y-DNA haplogroup
- E-V13>Z5018>FGC33625
- mtDNA haplogroup
- U1a1a
I haven't asked them anything. My questions will be about the "Kurdish" and whether I can get the Y positions tested in addition to the positive Y calls.
Cool, thanks for this. If they're rawdna is buggy i think its constructive if you let them know so they can nip it in the bud, anyway thanks so much for the valuable feedback, i'm following it closely : )
I1a3_Young
Regular Member
- Messages
- 556
- Reaction score
- 66
- Points
- 28
- Location
- FL
- Ethnic group
- Basically British
- Y-DNA haplogroup
- I1 Z63*
- mtDNA haplogroup
- H5b1
[TABLE="width: 337"]
[TR]
[TD][/TD]
[TD]Paper[/TD]
[TD]Ancestry[/TD]
[TD]LDNA[/TD]
[/TR]
[TR]
[TD]English[/TD]
[TD="align: right"]56.732%[/TD]
[TD="align: right"]69.0%[/TD]
[TD="align: right"]72.7%[/TD]
[/TR]
[TR]
[TD]Unknown, mostly brit[/TD]
[TD="align: right"]21.875%[/TD]
[TD][/TD]
[TD][/TD]
[/TR]
[TR]
[TD]Scots or Scots-Irish[/TD]
[TD="align: right"]7.837%[/TD]
[TD][/TD]
[TD="align: right"]8.6%[/TD]
[/TR]
[TR]
[TD]Norwegian[/TD]
[TD="align: right"]6.250%[/TD]
[TD="align: right"]10.0%[/TD]
[TD="align: right"]9.7%[/TD]
[/TR]
[TR]
[TD]German[/TD]
[TD="align: right"]3.906%[/TD]
[TD="align: right"]1.0%[/TD]
[TD][/TD]
[/TR]
[TR]
[TD]Irish[/TD]
[TD="align: right"]1.807%[/TD]
[TD="align: right"]10.0%[/TD]
[TD][/TD]
[/TR]
[TR]
[TD]Welsh[/TD]
[TD="align: right"]1.489%[/TD]
[TD][/TD]
[TD="align: right"]1.7%[/TD]
[/TR]
[TR]
[TD]French[/TD]
[TD="align: right"]0.098%[/TD]
[TD][/TD]
[TD][/TD]
[/TR]
[TR]
[TD]N. Italy[/TD]
[TD="align: right"]0.006%[/TD]
[TD="align: right"]3.0%[/TD]
[TD][/TD]
[/TR]
[TR]
[TD]Finland[/TD]
[TD][/TD]
[TD="align: right"]3.0%[/TD]
[TD="align: right"]3.1%[/TD]
[/TR]
[TR]
[TD]Basque[/TD]
[TD][/TD]
[TD="align: right"]3.0%[/TD]
[TD="align: right"]1.2%[/TD]
[/TR]
[TR]
[TD]Euro Jew[/TD]
[TD][/TD]
[TD="align: right"]1.0%[/TD]
[TD][/TD]
[/TR]
[TR]
[TD]"Kurdish"[/TD]
[TD] [/TD]
[TD] [/TD]
[TD="align: right"]3.0%[/TD]
[/TR]
[TR]
[TD][/TD]
[TD="align: right"]100.000%[/TD]
[TD="align: right"]100.000%[/TD]
[TD="align: right"]100.000%[/TD]
[/TR]
[/TABLE]
Here is a paper vs Ancestry vs LivingDNA breakdown. On LDNA I pulled Orkney into Scandinavian.
The unknown is all from the mother side with nothing non-British about it based on my mother's and her mother's Ancestry test.
[TR]
[TD][/TD]
[TD]Paper[/TD]
[TD]Ancestry[/TD]
[TD]LDNA[/TD]
[/TR]
[TR]
[TD]English[/TD]
[TD="align: right"]56.732%[/TD]
[TD="align: right"]69.0%[/TD]
[TD="align: right"]72.7%[/TD]
[/TR]
[TR]
[TD]Unknown, mostly brit[/TD]
[TD="align: right"]21.875%[/TD]
[TD][/TD]
[TD][/TD]
[/TR]
[TR]
[TD]Scots or Scots-Irish[/TD]
[TD="align: right"]7.837%[/TD]
[TD][/TD]
[TD="align: right"]8.6%[/TD]
[/TR]
[TR]
[TD]Norwegian[/TD]
[TD="align: right"]6.250%[/TD]
[TD="align: right"]10.0%[/TD]
[TD="align: right"]9.7%[/TD]
[/TR]
[TR]
[TD]German[/TD]
[TD="align: right"]3.906%[/TD]
[TD="align: right"]1.0%[/TD]
[TD][/TD]
[/TR]
[TR]
[TD]Irish[/TD]
[TD="align: right"]1.807%[/TD]
[TD="align: right"]10.0%[/TD]
[TD][/TD]
[/TR]
[TR]
[TD]Welsh[/TD]
[TD="align: right"]1.489%[/TD]
[TD][/TD]
[TD="align: right"]1.7%[/TD]
[/TR]
[TR]
[TD]French[/TD]
[TD="align: right"]0.098%[/TD]
[TD][/TD]
[TD][/TD]
[/TR]
[TR]
[TD]N. Italy[/TD]
[TD="align: right"]0.006%[/TD]
[TD="align: right"]3.0%[/TD]
[TD][/TD]
[/TR]
[TR]
[TD]Finland[/TD]
[TD][/TD]
[TD="align: right"]3.0%[/TD]
[TD="align: right"]3.1%[/TD]
[/TR]
[TR]
[TD]Basque[/TD]
[TD][/TD]
[TD="align: right"]3.0%[/TD]
[TD="align: right"]1.2%[/TD]
[/TR]
[TR]
[TD]Euro Jew[/TD]
[TD][/TD]
[TD="align: right"]1.0%[/TD]
[TD][/TD]
[/TR]
[TR]
[TD]"Kurdish"[/TD]
[TD] [/TD]
[TD] [/TD]
[TD="align: right"]3.0%[/TD]
[/TR]
[TR]
[TD][/TD]
[TD="align: right"]100.000%[/TD]
[TD="align: right"]100.000%[/TD]
[TD="align: right"]100.000%[/TD]
[/TR]
[/TABLE]
Here is a paper vs Ancestry vs LivingDNA breakdown. On LDNA I pulled Orkney into Scandinavian.
The unknown is all from the mother side with nothing non-British about it based on my mother's and her mother's Ancestry test.
I1a3_Young
Regular Member
- Messages
- 556
- Reaction score
- 66
- Points
- 28
- Location
- FL
- Ethnic group
- Basically British
- Y-DNA haplogroup
- I1 Z63*
- mtDNA haplogroup
- H5b1
I1a3_Young
Regular Member
- Messages
- 556
- Reaction score
- 66
- Points
- 28
- Location
- FL
- Ethnic group
- Basically British
- Y-DNA haplogroup
- I1 Z63*
- mtDNA haplogroup
- H5b1
[h=2]Note that the Kurdish only comes up when forced via "Complete" mode. It gives World (unassigned) in both Standard and Cautious
World (unassigned)[/h] When calculating your ancestry we've identified some DNA that is found in multiple areas of the world. These have been listed as "Unassigned", since we cannot pinpoint as confidently, specifically which regions that DNA is similar to. As our methods improve you will see your results updated and a lower proportion of Unassigned appearing.
I have seen a few more people with small % Kurdish.
I emailed them just now with the following demands, along with feedback:
1. Can I get my negative YSNP calls?
2. Can I repair/fill in/re-upload null values that I know from other tests such as Ancestry.com to get a more complete ethnicity estimate?
3. Can you give me details on the data and methods used to calculate the Basque and Kurdish?
I will keep the thread updated with response and response time.
World (unassigned)[/h] When calculating your ancestry we've identified some DNA that is found in multiple areas of the world. These have been listed as "Unassigned", since we cannot pinpoint as confidently, specifically which regions that DNA is similar to. As our methods improve you will see your results updated and a lower proportion of Unassigned appearing.
I have seen a few more people with small % Kurdish.
I emailed them just now with the following demands, along with feedback:
1. Can I get my negative YSNP calls?
2. Can I repair/fill in/re-upload null values that I know from other tests such as Ancestry.com to get a more complete ethnicity estimate?
3. Can you give me details on the data and methods used to calculate the Basque and Kurdish?
I will keep the thread updated with response and response time.
Johane Derite
Regular Member
- Messages
- 1,959
- Reaction score
- 1,057
- Points
- 113
- Ethnic group
- Albanian
- Y-DNA haplogroup
- E-V13>Z5018>FGC33625
- mtDNA haplogroup
- U1a1a
I know im becoming repetitive but thanks for the updates im really interested in them
I1a3_Young
Regular Member
- Messages
- 556
- Reaction score
- 66
- Points
- 28
- Location
- FL
- Ethnic group
- Basically British
- Y-DNA haplogroup
- I1 Z63*
- mtDNA haplogroup
- H5b1
Response received today.
1. Cannot have YSNP negative calls because the file would be too large (16,000 calls) and they don't have a nice way to present it.
2. Cannot accept any 2nd party data for consistency purposes (obvious, I just thought I'd ask on a long shot)
3. Generic response about how they calculate ethnicity but no specifics for the Kurdish or Basque provided.
1. Cannot have YSNP negative calls because the file would be too large (16,000 calls) and they don't have a nice way to present it.
2. Cannot accept any 2nd party data for consistency purposes (obvious, I just thought I'd ask on a long shot)
3. Generic response about how they calculate ethnicity but no specifics for the Kurdish or Basque provided.
I1a3_Young
Regular Member
- Messages
- 556
- Reaction score
- 66
- Points
- 28
- Location
- FL
- Ethnic group
- Basically British
- Y-DNA haplogroup
- I1 Z63*
- mtDNA haplogroup
- H5b1
There are 150,540 SNPs common to both the LDNA and Ancestry files that GEDmatch uses. When comparing Ancestry files they use 439,216 SNPs. This is probably why puntDAL is terrible with LivingDNA data - it's looking for SNPs that aren't there.
LDNA plays well with GEDmatch Eurogenes and MDLP but not with puntDNAL.
Jtest with LDNA file shows less than 1% Ashkenazi but Oracle calls English/English/English/English. Ancestry test Oracles English/English/English/W. German
I ran all kits for both side of my family and all of them show about the same Ash% on Jtest. I can't find any specific sources on how the mystery near-east or unassigned is determined by any calculator.
LDNA plays well with GEDmatch Eurogenes and MDLP but not with puntDNAL.
Jtest with LDNA file shows less than 1% Ashkenazi but Oracle calls English/English/English/English. Ancestry test Oracles English/English/English/W. German
I ran all kits for both side of my family and all of them show about the same Ash% on Jtest. I can't find any specific sources on how the mystery near-east or unassigned is determined by any calculator.
Last edited:
I1a3_Young
Regular Member
- Messages
- 556
- Reaction score
- 66
- Points
- 28
- Location
- FL
- Ethnic group
- Basically British
- Y-DNA haplogroup
- I1 Z63*
- mtDNA haplogroup
- H5b1
srdceleva
Banned
- Messages
- 414
- Reaction score
- 76
- Points
- 0
- Location
- Austria
- Ethnic group
- 75% Slovak, 25℅ American mix
- Y-DNA haplogroup
- R1a-m458(L260)
- mtDNA haplogroup
- U4b1b1
Is living DNA still working to update their raw data? Does anyone know this or are we just hoping gedmatch will eventually be able to process living DNA kits better. Even with Genesis it only takes a couple of seconds for each calc to read my living DNa data. The snp countmust be extremely low.There are 150,540 SNPs common to both the LDNA and Ancestry files that GEDmatch uses. When comparing Ancestry files they use 439,216 SNPs. This is probably why puntDAL is terrible with LivingDNA data - it's looking for SNPs that aren't there.
LDNA plays well with GEDmatch Eurogenes and MDLP but not with puntDNAL.
Jtest with LDNA file shows less than 1% Ashkenazi but Oracle calls English/English/English/English. Ancestry test Oracles English/English/English/W. German
I ran all kits for both side of my family and all of them show about the same Ash% on Jtest. I can't find any specific sources on how the mystery near-east or unassigned is determined by any calculator.
Sent from my KIW-L21 using Tapatalk
I1a3_Young
Regular Member
- Messages
- 556
- Reaction score
- 66
- Points
- 28
- Location
- FL
- Ethnic group
- Basically British
- Y-DNA haplogroup
- I1 Z63*
- mtDNA haplogroup
- H5b1
LDNA uses a newer chip for their tests. They can't change which SNPs are included. The GEDmatch calcs are the ones that need to be updated to use more SNPs.Is living DNA still working to update their raw data? Does anyone know this or are we just hoping gedmatch will eventually be able to process living DNA kits better. Even with Genesis it only takes a couple of seconds for each calc to read my living DNa data. The snp countmust be extremely low.
Sent from my KIW-L21 using Tapatalk
GEDmatch Genesis calcs give me more specific results on the oracles.