real expert
Regular Member
- Messages
- 795
- Reaction score
- 455
- Points
- 63
Abstract
Arab populations are largely understudied, notably their genetic structure and history. Here we present an in-depth analysis of 6,218 whole genomes from Qatar, revealing extensive diversity as well as genetic ancestries representing the main founding Arab genealogical lineages of Qahtanite (Peninsular Arabs) and Adnanite (General Arabs and West Eurasian Arabs). We find that Peninsular Arabs are the closest relatives of ancient hunter-gatherers and Neolithic farmers from the Levant, and that founder Arab populations experienced multiple splitting events 12–20 kya, consistent with the aridification of Arabia and farming in the Levant, giving rise to settler and nomadic communities. In terms of recent genetic flow, we show that these ancestries contributed significantly to European, South Asian as well as South American populations, likely as a result of Islamic expansion over the past 1400 years. Notably, we characterize a large cohort of men with the ChrY J1a2b haplogroup (n = 1,491), identifying 29 unique sub-haplogroups. Finally, we leverage genotype novelty to build a reference panel of 12,432 haplotypes, demonstrating improved genotype imputation for both rare and common alleles in Arabs and the wider Middle East.
Fig. 1: Genetic structure of the QGP population.
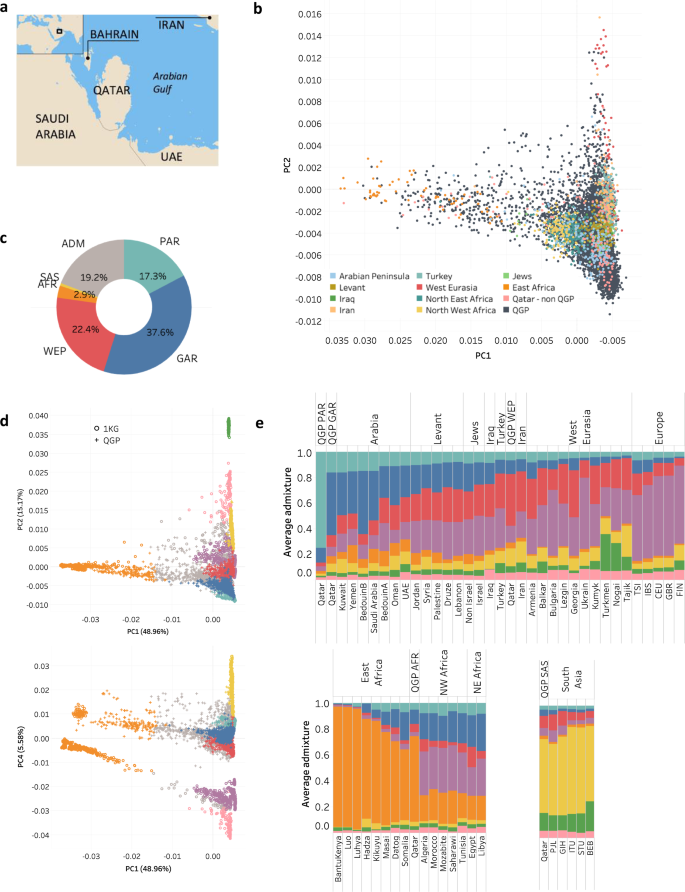
a Map showing the geographical location of Qatar, source of the study population. b Principal Component Analysis plot showing overlap of Qatar Genome Program (QGP) subjects with populations from the wider Middle Eastern region found in the Human Origin, Greater Middle East and other public datasets. QGP samples are shown in black and other reference populations in various colors. c Genetic sub-groups of the Qatari population based on dominant ancestral fraction (≥0.5) and k = 8. The abbreviations refer to Peninsular Arabs (PAR), General Arabs (GAR), Arabs of West Eurasia and Persia (WEP), South Asian Arabs (SAS), African Arabs (AFR), Admixed Arabs (ADM). d PCA showing QGP sub-groups in the context of continental populations form Africa, Europe, South Asia, East Asia and America. e Average ancestry fractions for QGP and other world populations (k = 8). The three sub-panels highlight various reference populations as relevant to the QGP subpopulations. Colors in panels (c–e) are the same as those used to delineate the distinct ancestral fractions in ADMIXTURE. Abbreviations of the 1KG subpopulations are: BEB Bengali from Bangladesh, CEU Utah Residents (CEPH) with Northern and Western European Ancestry, FIN Finnish in Finland, GBR British in England and Scotland, GIH Gujarati Indian from Houston, Texas, IBS Iberian Population in Spain, ITU Indian Telugu from the UK, PJL Punjabi from Lahore, STU Sri Lankan Tamil from the UK, TSI Toscani in Italy. Source data are provided as a Source Data file.
https://www.nature.com/articles/s41467-021-25287-y
Arab populations are largely understudied, notably their genetic structure and history. Here we present an in-depth analysis of 6,218 whole genomes from Qatar, revealing extensive diversity as well as genetic ancestries representing the main founding Arab genealogical lineages of Qahtanite (Peninsular Arabs) and Adnanite (General Arabs and West Eurasian Arabs). We find that Peninsular Arabs are the closest relatives of ancient hunter-gatherers and Neolithic farmers from the Levant, and that founder Arab populations experienced multiple splitting events 12–20 kya, consistent with the aridification of Arabia and farming in the Levant, giving rise to settler and nomadic communities. In terms of recent genetic flow, we show that these ancestries contributed significantly to European, South Asian as well as South American populations, likely as a result of Islamic expansion over the past 1400 years. Notably, we characterize a large cohort of men with the ChrY J1a2b haplogroup (n = 1,491), identifying 29 unique sub-haplogroups. Finally, we leverage genotype novelty to build a reference panel of 12,432 haplotypes, demonstrating improved genotype imputation for both rare and common alleles in Arabs and the wider Middle East.
Fig. 1: Genetic structure of the QGP population.

a Map showing the geographical location of Qatar, source of the study population. b Principal Component Analysis plot showing overlap of Qatar Genome Program (QGP) subjects with populations from the wider Middle Eastern region found in the Human Origin, Greater Middle East and other public datasets. QGP samples are shown in black and other reference populations in various colors. c Genetic sub-groups of the Qatari population based on dominant ancestral fraction (≥0.5) and k = 8. The abbreviations refer to Peninsular Arabs (PAR), General Arabs (GAR), Arabs of West Eurasia and Persia (WEP), South Asian Arabs (SAS), African Arabs (AFR), Admixed Arabs (ADM). d PCA showing QGP sub-groups in the context of continental populations form Africa, Europe, South Asia, East Asia and America. e Average ancestry fractions for QGP and other world populations (k = 8). The three sub-panels highlight various reference populations as relevant to the QGP subpopulations. Colors in panels (c–e) are the same as those used to delineate the distinct ancestral fractions in ADMIXTURE. Abbreviations of the 1KG subpopulations are: BEB Bengali from Bangladesh, CEU Utah Residents (CEPH) with Northern and Western European Ancestry, FIN Finnish in Finland, GBR British in England and Scotland, GIH Gujarati Indian from Houston, Texas, IBS Iberian Population in Spain, ITU Indian Telugu from the UK, PJL Punjabi from Lahore, STU Sri Lankan Tamil from the UK, TSI Toscani in Italy. Source data are provided as a Source Data file.
Comparison to Middle Eastern populations and nature of QGP ancestries
To determine the nature of the identified QGP ancestry clusters, notably cyan, blue and red, we examined their co-localization on PCA and Admixture sharing relative to publicly available samples from diverse Middle Eastern populations6,10,13,14,29. As highlighted in Supplementary Fig. 11, the Blue cluster overlapped largely with samples from Arabia, Levant (including both Arab and Jewish populations) and North Africa. The Red cluster predominantly overlapped with Persians, Turkish and other West Eurasians. Interestingly the Cyan cluster did not overlap with public samples except previously published Qataris14 (Supplementary Fig. 11). Consistently, patterns of admixture fractions reflect these relationships and show a number of gradients on the axis from Arabia to Europe (Fig. 1e): (1) Decrease of Cyan and Blue signatures (2) Increase of Purple (dominant ancestry in Europeans) (3) Increase of Red signature towards West Eurasia followed by a decrease towards Europe. We note that for Levant populations, Jewish and Arab populations have similar Admixture patterns reflecting their common ancestral history. The QGP Orange and Yellow clusters have similar signatures to other Eastern African and South Asian populations, respectively. For privacy reasons, it was not possible to get the tribal affiliations of QGP samples, however based on aggregate information we could trace the Cyan cluster to tribes originating from South of Arabia, Blue to the Levant/North Africa and Red to Persia. Therefore, based on these analyses, we name the QGP clusters as: Blue: General Arabs (GAR), Cyan: Peninsular Arabs (PAR), Red: Western Eurasian and Persian Arabs (WEP), Yellow: South Asian Arabs (SAS), Orange: African Arabs (AFR) and Gray: Admixed Arabs (ADM). This further refines the previously described breakdown of the Qatari population into three groups of Bedouins (Q1), Persian/South Asians (Q2) and Africans (Q3)9,12. We note that most public samples from Arabia used in our comparison including Bedouins, cluster closer to GAR than to PAR, which suggests their origin in Levant/North Arabia.
Discussion
This study provides an in-depth analysis of the genomic structure of the QGP phase 1 cohort, representing a comprehensive set of genomes from a Middle Eastern population. Despite the relatively small size of Qatar, it revealed unique and shared ancestries reflecting the wider Middle Eastern region and its centrality to recent and distant history.
The large dataset allowed the refinement of previously described genetic ancestries in the Qatari population (Q1, Q2 and Q3)12, identifying five main distinct ancestries that are generalizable to Arab and other Middle Eastern populations (PAR, GAR, WEP, SAS, AFR). PAR, GAR and WEP were found to be descendants of the main Arab genealogical branches of Qahtanite and Adnanite, which refer to indigenous Arab and Arabized populations respectively. The terms ‘Arab Bedouin’ and ‘indigenous Arab’ were used interchangeably in previous studies, however we find that in many cases this is inaccurate. Modern gene flow was identified from these ancestries to various European, South Asian and South American populations, likely reflecting post-Islamic expansion.
We performed an estimation of time divergence of Arab populations using hundreds of genomes. This pointed to an ancestral Arab population that dominated the Levant region, and split into modern lineages around 12–20 kya. Analysis involving ancient human DNA dating back to various archeological periods indicated that Peninsular Arabs are ancestral to modern Middle Eastern ancestries, being the closest to the basal founders that populated the ancient Near East. The sequencing of ancient genomes from the Arabian Peninsula will shed more light on the contribution of native Arabs to early out-of-Africa migrations.
In addition to enhancing the resolution on population structure, we performed ROH analysis using whole-genome data for a Middle Eastern population and at large scale, identifying some of the highest levels of human ROH ever reported to date. This highlights the tribal, endogamous nature of Arab society and culture, as well as the utility of next-generation sequencing to uncover recessive founder pathogenic alleles in high consanguinity setting8. Population-specific boundaries defining various ROH classes were calculated, showing an upper shift in Arab in comparison to world populations. Arabs also exhibited the least short ROH after African populations, reflecting their proximity to out-of-Africa migrations.
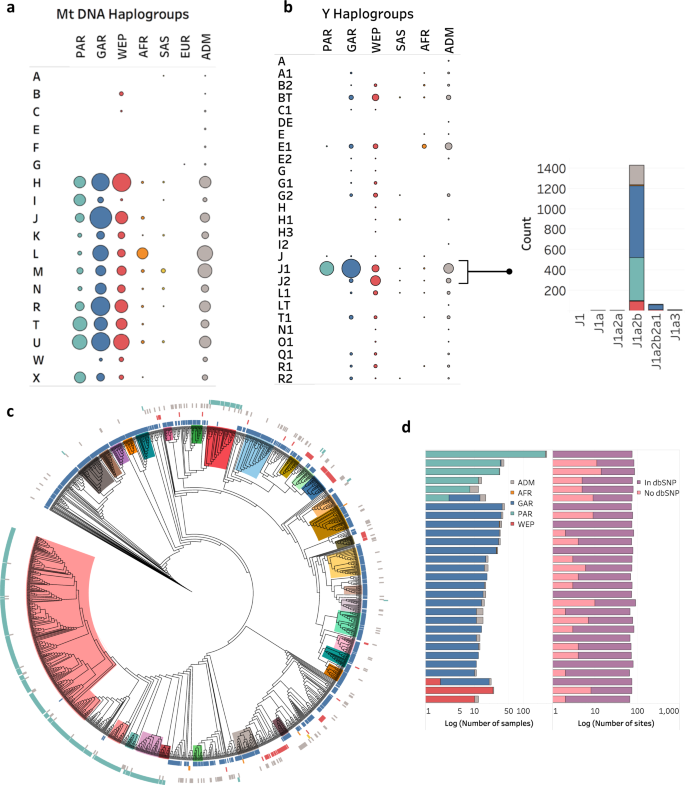
Furthermore, we report a substantial number of sequenced individuals having the J1a2b haplogroup which we further characterize into 29 novel sub-haplogroups. These sub-haplogroups were found to partition well among the autosomal ancestries, reinforcing the tribal, patrilocal nature of the regional populations.
Finally, a dedicated QGP imputation panel was generated to leverage this dataset, which shall complement the currently available panels by providing more accurate imputation of Arab and Middle Eastern genomes. This will enable association studies with greater scale and statistical power to detect causal variants underlying biological traits and diseases. Notably, there is an ongoing effort to build a population genotypic array (Q-chip) that would leverage rare and novel missense/LoF variants in disease genes from this panel (Rodriguez-Flores et al. In press). The current dataset has recently helped identify novel loci associated with a range of clinically relevant traits60,61. The value of this resource will expand in the upcoming phases of QGP as tens of thousands of additional subjects will be sequenced over the next few years, enabling vital future genomic and medical research in the Middle East and globally.
https://www.nature.com/articles/s41467-021-25287-y