Introduction
A long standing traditional cultural division exists between the Irish, the Welsh and the Highland Scots on one side, who are of Celtic heritage, and the English on the other side, who are of mixed Germanic, French, Celtic and even Roman ancestry. Until recently, no historian could really agree on much influence the Romans, Anglo-Saxons, Vikings and Normans had had on the British and Irish gene pools, let alone agree on where the Celts came from before that. Were the Celts descended from Bronze Age or the Iron Age invaders from the Continent? Or perhaps were they even the direct descendants of the Neolithic farmers who built Stonehenge, or even of the Mesolithic hunter-gatherers who recolonised northern Europe at the end of the last glaciation 10,000 years ago? Before the age of historical population genetics these questions were left to anyone's guess, and indeed even academics fought bitterly over such speculations. The fast paced progress in genetic sequencing have shed some new light on our prehistory and it is now possible to answer all those questions with a great degree of certainty. This page attempts to retrace the origins of the British and Irish people through the analyses of their genetic markers using Y-chromosomal (Y-DNA) haplogroups, which are passed on nearly unaltered from father to son, mitochondrial DNA (mtDNA), which is inherited only from one's mother, and complete genomic studies looking at all the chromosomes.
Genetic history of the British Isles
| If you are new to population genetics... |
|---|
|
In the following section we will review the Y-DNA haplogroups of the various prehistoric and historical populations that have settled in Great Britain and Ireland since Cro-Magnon colonised Europe during the Ice Age. that If you are unfamiliar with haplogroups or population genetics, we recommend that you familiarise yourself first with the basics by viewing the Video Tutorials about genetics and read our Frequently Asked Questions about DNA tests. Each haplogroup corresponds to a distinct ancestral lineage. Haplogroups are divided into numerous levels of subclades that form a phylogenetic tree, which is just a fancy word for genealogical tree of genetic ancestry. You may also find it useful to visualise the modern distribution of Y-DNA haplogroups to get a sense of they represent. Detailed descriptions of each haplogroup and their history are available here, but links to each haplogroup's page are also provided in the text below.
|
Paleolithic to Early Neolithic
During the Paleolithic period, the British Isles were covered by a thick sheet of ice, which made them all but uninhabitable. Europe had been originally inhabited by Neanderthals, a close relative of anatomically modern humans (Homo sapiens). Neanderthals had been living in Ice Age Europe for over 250,000 years when Homo sapiens started to advanced into Europe from the Middle East from 45,000 years ago, reaching western Europe approximately 35,000 years ago. The last pure Neanderthals may have survived until 24,000 years ago around Gibraltar. The skull of a 4 year-old Neanderthal boy displayed signs of hydridisation bewteen Neanderthals and Homo sapiens. It is now known that all modern Europeans and Asians carry a few percent's of Neanderthalian DNA due to such hybridisation.
Only one Paleolithic Y-DNA has beentested (by Seguin-Orlando et al. 2014 to date, a 37,000 year-old individual from southern Russia, and belonged to Y-haplogroup C1 and mt-haplogroup U2. Haplogroup C1 (actually C1a2) also shows up in a sample from Mesolithic Spain 7,000 years ago (Sánchez-Quinto et al. 2012) - a surprising continuity in both time and space.
It likely that the first migrations of Homo sapiens to Europe, linked with the development of Aurignacian culture, brought old haplogroups such as C-V20 and H-P96, simply because other haplogroups further down the phylogenetic tree did not exist yet. It is possible that even the West African haplogroup A1a reached Europe during the Paleolithic by crossing the Strait of Gibraltar, as some sub-Saharan African admixture regularly shows up in Mesolithic and Paleolithic samples, and modern A1a samples has been found in the northern fringes of Europe (Ireland, Norwegian Lappland, Finland) which cannot be explained by any more recent migration.
The robust people of the Aurignacian were succeeded by more gracile people during the Gravettian period (32,000-22,000 years ago). This second wave of modern humans is probably the one that brought haplogroup I. A third wave in the late glacial period (17,000-12,000 years ago) might have brought E-M78 (V12, V13 and V22 subclades) from North Africa to Iberia, Italy and Greece, while J2b expanded from Anatolia to the southern Balkans. Haplogroup R1a would have arrived from Siberia to eastern Europe following the end of the last glaciation.
During the Last Glacial Maximum (LGM), which lasted approximately from 26,500 to 19,000 years ago, the glaciers descended all the way to the Alps and Cro-Magnons sought refuge in temperate regions of southern Europe, then re-expanded north once the climate warmed up and the ice melted. The few Mesolithic hunter-gatherers tested to date belonged to Y-haplogroups C1, F and I2.
Whereas Y-haplogroups C1 and F are almost extinct in Europe today, there are a few surviving Cro-Magnon paternal lineages in the British Isles today, mostly haplogroup I2a1 and I2a2a, who may well be direct descendants from those Mesolithic hunter-gatherers that first set foot in the British Isles after the glaciers retreated. Other Mesolithic lineages survived in central Europe and Scandinavia and would have come to the British Isles during the Iron Age with the Celts (I2a2b) and Germanic peoples (I1, I2a2a-Z161).
Ancient DNA tests have confirmed haplogroup U5 as the dominant female European lineage during the Mesolithic, although a few U2e and U4 samples were also found in Northeast Europe. Nowadays U5 is found in 8.5% of the British and Irish population, although not all of it descends from the Mesolithic inhabitants of the Isles. Since U5 is the only haplogroup that has been consistenly present through the ages since the Mesolithic in all western, central and northern Europe, a substantial part of modern U5 lineages could have come from any of the peoples who migrated to Britain and Ireland in the last 8,000 years.
Neolithic newcomers
The Neolithic period saw the advent of agriculture (livestock herding, then cereal farming), the use of pottery and the replacement of the nomadic lifestyle by sedentism. Originating in the Fertile Crescent circa 9500 BCE, the Neolithic spread through Anatolia and Greece, then its diffusion split in two routes. The southern route went across the Mediterranean to Italy, southern France and Iberia, and evolved into the Printed-Cardium Pottery culture (5000-1500 BCE). The northern branch developed in the Balkans as the Starčevo culture (6000-4500 BCE) and the Linear Pottery culture (LBK) (5500-4500 BCE), following the Danube and its tributaries until Germany, northern France and the Benelux. It has been suggested that the modern divide between northern European cultures, favouring butter-based cooking, and southern European cultures, preferring olive oil-based cuisine, dates back from the Neolithic period.
Neolithic herders and farmers from the Near East appear to have belonged predominantly to haplogroup G2a, with minorities of J1 and T. Ancient DNA tests have confirmed the present of G2a in all Neolithic sites in Europe tested to date, including numerous samples from the Cardium Pottery, Starčevo and and Linear Pottery cultures. Nevertheless, substantial minorities of other haplogroups have been found on different Neolithic sites next to a G2a majority, including C1a2, F, H2, I*, I1, I2a and I2a1 in Central and Southeast Europe (LBK, Starčevo) and E-V13 and I2a in Spain and France (Cardium Pottery). Many of these (except H2) probably represent assimilated hunter-gatherers descended from Mesolithic and Paleolithic Europeans.
R1b-V88 cattle herders from the Near East also managed to get to Western Europe via North Africa, probably crossing at Gibraltar. The oldest known R1b-V88 sample in Europe is a 7,100 year-old from the Catalan Pyrenees tested by Haak et al. 2015 and whose autosomal DNA matched that of other Neolithic farmers, without a trace of Steppe ancestry as in Bronze Age R1b-L11 samples (see below). Nowadays R1b-V88 only represent a tiny fraction of European R1b. Even in Ireland, where R1b exceeds 80% of the population, only about 1% belong to the Neolithic North African V88 subclade.
Maternal lineages brought by Neolithic farmers from the Balkans and Anatolia can be safely ascertained by the large number of ancient mtDNA already tested. They included haplogroups HV, H2a2, H5a, H13, H20, J1c, J2b1a, K1a, N1a, T2 and X.
There is overwhelming evidence that Neolithic farmers intermingle with some of the Mesolithic foragers they encountered. Haplogroups C1a, F, I1 and I2 were all found next to G2a in various sites tested. Additionally, the Sardinians are the modern European population who are the most closely related to Neolithic farmers, and 37.5% of the male population belongs to Y-haplogroup I2a1a (CTS595). I2a lineages were found alongside G2a among early Neolithic farmers in Serbia and in Languedoc (southern France), close to the Spanish border. Nowadays the second highest frequency of I2a1a is found among the Basques, but this paternal lineage is found all over western Europe, probably as a remnant of the Megalithic culture that unified Atlantic and the West Mediterranean regions of Europe throughout the Neolithic and Chalcolithic periods (c. 4000-2000 BCE in Britain). The ancient Y-DNA samples from Atlantic Megalithic cultures tested to date included haplogroups G2a (20 samples), H2, I2a1 (4 samples) and I2a2a.
In other words, Megalithic structures like Stonehenge in Wiltshire, the Newgrange passage tomb in County Meath, Ireland, or the Callanish Stones on the Isle of Lewis in Scotland were in all likelihood built by men belonging essentially to haplogroups G2a and I2a.
Megalithic mtDNA from Brittany is a typical blend of Mesolithic (H1, H3, U4, U5b, V) and Neolithic (H5, K1a, N1a, T2, X2) lineages, in direct continuity of the Cardium Pottery and Linear Pottery cultures. Consequently, Megalithic people were predominantly G2a and I2a people, probably with minorities of E-M78, R1b-V88 and T1a in southern Europe.
Late Neolithic to Early Bronze Age
The Late Neolithic period saw the development of the Bell Beaker phenomenon. Often considered erroneously as a single culture, it was almost certainly not a unified entity, be it politically, linguistically or ethnically, but rather a vast multicultural trade network. For the next 500 years it would spread on land and through maritime routes to various isolated regions in western Europe, including most of England, Wales, Lowland Scotland and parts of Ireland, but also to the Brittany, Portugal, several regions of Spain, the Low Countries, Jutland, southern Germany, the Rhône valley, the Alps, northern Italy, Sardinia, and as far east as Bohemia. Most of these regions (except central Europe) were already somehwat linked to each others as members of the Megalithic culture, which evolved from the Early Neolithic cultures.
The Bell Beaker cultural phenomenon did not in fact replace the Megalithic culture in western Europe, but coincided with it. The Beaker people continued to use common Megalithic burials (e.g. passage graves) like their Neolithic ancestors. In central Europe, where no Megalithic culture existed, bell beaker artefacts nevertheless appear due to the presence of western European merchants.
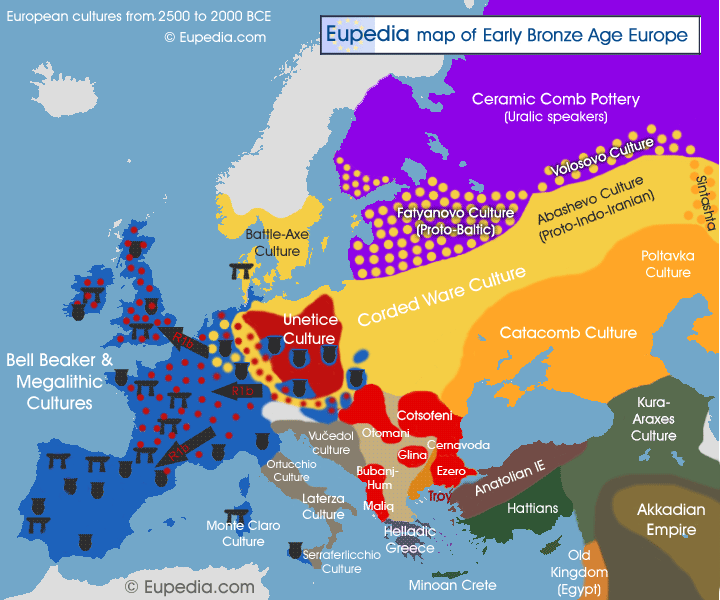
It is perhaps the wealth of Megalithic people that attracted, through the Beaker network, the Indo-European speakers from central Europe, and caused them to invade western Europe and destroy the Megalithic cultures that had lasted for several millennia. Equipped with bronze weapons and horses, these Indo-Europeans were not cereal farmers but cattle herders from the Pontic Steppe, north of the Black Sea. They had already conquered the Balkans, the Carpathians, Poland, Germany, Scandinavia and the Baltic countries between 4,000 and 2,800 BCE, causing the collapse of all the Chalcolithic and Neolithic cultures in those areas. The southern R1b branch had advanced from the Hungarian plain to Bohemia and Germany by 2500 BCE (presence of R1b confirmed by Lee at al. 2012), and continued its migration until the Atlantic coast, reaching Britain and western France by 2,200 BCE and Ireland by 2,000 BCE.
It is likely that these Proto-Celts who invaded the British Isles belonged to a great majority to the L21 subclade of R1b, as this haplogroup now makes up over two thirds of paternal lineages in Wales, Ireland and Highland Scotland. In fact, Cassidy et al. (2015) tested the genomes of three Bronze Age individuals from Rathlin Island in Ireland dating from between 1550 and 2000 BCE, and found out that they all belonged to R1b-L21 and were almost genetically indistinguishable from the genomes from the Unetice culture in Central Europe at the same period, confirming that a migration from central Europe to Ireland had brought R1b-L21 by 2,000 BCE. In contrast, Cassidy's team found that the genomes of the Neolithic inhabitants of Ireland were almost identical to those of other Neolithic farmers in Europe and did not carry Steppe ancestry.
If we exclude haplogroups of Germanic origins (Anglo-Saxon, Viking, Norman), it can be estimated that over 90% of British and Irish men carried the R1b-L21 Y chromosome by the end of the Bronze Age. This makes R1b-L21 the quintessential Gaelic paternal lineage. It is not clear how such a drastic change in paternal lineages happened, especially since maternal lineages were much less affected by those Celtic migrations. It has been postulated that technologically superior Proto-Celtic warriors, equipped with Bronze weapons and riding on horses, could have massacred or enslaved indigenous men while taking their women, or that they established a ruling elite that passed on more Y chromosomes through sustained polygamy over many centuries.
Late Bronze Age to Iron Age
Alpine Celts
A latter Celtic migration took place around 500 BCE, when Central European Celts from the Hallstatt culture expanded over a large swathe of western Europe. Travelling with their families on wagons transporting their belongings, the Celts colonised most of France, Belgium and the south-east of England. Soon afterwards the classical Iron Age Celtic La Tène culture flourished around the Alps and quickly overran Gaul, northern Italy and a big part of central Europe. In the first century BCE, several La Tène tribes like the the Belgic Atrebates and the Gaulish Parisii crossed over the Channel and settled in southern England and Yorkshire. The cultural influence of La Tène spread to the whole of Ireland in the 1st century BCE. It is possible that the Picts, whose art was characteristic of the La Tène style, descended from undocumented La Tène Celts, or perhaps from an offshoot of the Parisii from Yorkshire displaced northward by the Romans.
No ancient DNA from the Hallstatt and La Tène culture has been tested to date, but it is very likely that these Celts carried a large percentage of R1b-U152 (mostly the L2 subclade), and probably low percentages of G2a-U1, I2a2b (L38) and J2b2 lineages, among others. R1b-U152 is found at low frequency almost everywhere in the British Isles, but is considerably more common in eastern and southern England (5-10%), reaching a peak of nearly 15% in East Anglia and in Kent.

Another lineages associated with Celtic people is R1b-DF27. This haplogroup is found primarily in the Iberian peninsula and western France, but makes up between 4 and 12% of the male lineages in Britain and Ireland today, except in the Scottish Highlands where its is for the most part absent. It is not yet clear when and how DF27 reached the archipelago, but its higher frequency in southern and central England and its declining frequency toward western Wales and the Scottish Highlands suggests that it would have come from the continent during the Atlantic Bronze Age (1300-700 BCE) and/or with subsequent Hallstatt, La Tène and Belgic migrations. It is also possible that a minority of DF27 lineages came from Gaul and Iberia during the Roman period, when migration within the empire was easy. The very low frequency of DF27 in northern Germany and Scandinavia today rules out a Germanic origin for the greatest part of DF27 lineages in Britain and Ireland today. The Normans might have brought more DF27 - among the numerous other French and Scandinavian lineages that could have been found among the Norman conquerors.
No proper study of deep mtDNA subclades has been conducted yet, but a rough estimation is that between 20% and 35% of maternal lineages can be traced back to Indo-European invaders, be them Celtic, Roman or Germanic. The disparity between paternal and maternal Indo-European lineages is not surprising considering that Proto-Indo-European speakers advanced across Europe, from the Black Sea shores, as military conquerors, and progressively blended with the conquered populations by taking local wives or concubines. Consequently, whereas their paternal lineages spread like a wild fire in all conquered regions, we observe a slowly declining gradient from east to west for maternal haplogroups.
Romans
In 43 CE, under the reign of Emperor Claudius, the Romans invaded Great Britain. Within two decades most of England and Wales had been conquered and would remain a province of the Roman Empire for over three centuries. For most of the second century Lowland Scotland was also Romanised. The Romans established over 70 cities or towns in Britain, including London, St Albans, Colchester, Winchester, Gloucester, Exeter, Leicester, Lincoln, Manchester and York.
It is very difficult to assess the genetic impact of Romanisation on the British population as the Roman citizens, soldiers and slaves who settled in Britannia were not merely people from the Latium or Italy, but could have come from anywhere in the empire. Even if we assume that Britain was fully Celtic before the Roman conquest, similar to Ireland or the Scottish Highland once Germanic DNA has been removed, it is still very hard at present to clearly differentiate Brythonic Celts from other Celts from the continent, notably Gaul, who might have settled in Roman Britain. Even the Romans from Italy appear to have belonged predominantly to the same R1b-U152 as Hallstatt and La Tène Celts, also accompanied by significant minorities of G2a-U1 and J2b. Deeper subclade analysis may soon allow population geneticists to distinguish between Roman/Italic and Celtic subclades within these haplogroups. At present it seems that the L2 and Z36 subclades of R1b-U152 are more Celtic/Gaulish, while Z56 and Z192 are more Italic/Roman, but it may be more complicated than that.
Overall, the majority of immigrants to Roman Britain came from the Mediterranean region, with higher percentages from Italy and the Balkans reported for Roman legions settled in Britain. There may also have been Near Eastern merchants, like the Jews, whose diaspora started soon after the Roman conquest of Britain. Ancient Mediterranean people would have carried mostly haplogroups such as E1b1b, J1, J2a, and to a lower extent also G2a and T. Nowadays, those haplogroups are considerably rarer among the Irish and Highland Scots, and inexistent in remote islands like Orkney or Shetland (except for haplogroup T). Even in Ireland such lineages are typically found among people with English or Norman surnames. Wales, which supposedly served as a refuge to the Romano-British population fleeing the Anglo-Saxon invasions, has by far the highest percentages of haplogroup E1b1b, G2a and T (total 7.5% against 2.5% for Scotland and 3% for Ireland), while England has the highest percentage of J2 (3.5%).
In 2014, Schiffels and colleagues sequenced whole genomes from five skeletons excavated in Hinxton, Cambridgeshire, including two of Iron Age Britons (the other three were from early medieval Anglo-Saxons). The analysis of the Hinxton genomes revealed that pre-Roman Celtic Britons did not have any West Asian or Southwest Asian genetic admixture in them. In contrast, modern Britons possess respectively 6.7% and 1.2%. The Anglo-Saxon samples had non-null but negligible levels (1.70% and 0.8%) comparable to modern Finnish or Icelandic people.
This suggests that the West Asian or Southwest Asian admixtures entered the British gene pool during the Roman and/or Norman period, since neither the Anglo-Saxons nor the Vikings could have contributed to such levels. The French only have very slightly higher levels of each admixture than the British, and the Normans actually have comparable levels, so the only way they could have been responsible for the modern British admixtures would be through nearly total population replacement, which is utterly unthinkable.
The Italians have considerably higher levels of West Asian or Southwest Asian admixtures. Modern South Italians have 25% and 9% respectively, Central Italians have 22% and 7%, and even the highly Celticised and Germanised North Italians, who boast higher percentages of R1b than the British, still have 13% and 3%. Unfortunately no ancient Italian genome has been tested yet, but it is doubtful that the ancient Romans possessed less West Asian or Southwest Asian DNA than modern Italians since the only major migrations to Italy since then were those of Germanic peoples (Goths, Vandals, Lombards, Franks, Normans) who would have diluted such admixtures.
Considering that ancient Rome was a huge city and had a very cosmopolitan population with immigrants from all over the empire, and that the population of the city dispersed into the surrounding countryside in the early Middle Ages, we could assume that Central Italians are fairly representative of the average Roman Empire citizenry and therefore could be used as a proxy for the genetic admixture of the various immigrants who settled in Roman Britain. Obviously Central Italians have a higher proportion of Italian blood than from immigrants from other regions of the empire, but that must also have been the case for Roman Britain. The majority of administrators, land owners and legionaries (at least in the first century of Romanisation) would have hailed from Italy. Using Central Italians as a proxy, and hoping that the Hinxton samples are representative of pre-Roman Britons, it is possible that approximately one third of the autosomal genes in the British population comes from Mediterranean people who settled in Britain during the Roman period, but also via the Normans, who had substantial French admixture. The French having higher Neolithic and Roman ancestry than Iron Age Britons, the Normans would have indubitably raised the share of Mediterranean ancestry in the British Isles.
How does that compare to Y-chromosomal DNA? There is no way of knowing for sure how much of the British R1b-U152 is Alpine Celtic, how much is Roman, and in both cases how much came with the Normans. Likewise G2a and even maybe E-V13 and T could have come from Neolithic farmers, continental Celts, Romans or even Germanic people. Making the extreme hypothetical assumption that Neolithic paternal lineages had been entirely replaced by Bronze and Iron Age Celts, that Britain was purely R1b before the Roman period, and that Germanic migrations only brought R1b, R1a, I1, I2a2 and Q, and ignoring completely Norman contributions to the Y-chromosomal structure of the British population, this results in a maximum roughly 15% of male lineages of "Roman" origin for England, 10% for Wales, and 7% for Scotland. However, it would be reasonable to assume that at least half of these come from Alpine Celts and Normans, and probably more in Scotland's case. It is hard to explain the discrepency with the 30-35% of autosomal genes of Mediterranean origin. One explanation is that a substantial share of Romano-British male population was killed by the invading Anglo-Saxons, and that autosomal genes were passed on through Roman-British mothers who bore the children of the Germanic invaders (a similar scenario to the Proto-Celtic invasions, but with a less extreme male population replacement).
Anglo-Saxons & Vikings
from 410 to the middle of the 6th century, the Jutes, the Angles and the Saxons, Germanic tribes from Denmark and north-west Germany, left their homeland and migrated to what would become England. The Jutes settled in Kent and Hampshire, the Saxons in the rest of southern England as far north as the Worcestershire, while the Angles occupied the rest of England from East Anglia to Northumberland and southeastern Scotland.
A few centuries later, from around 800, Danish Vikings started raiding the coastlines of the British Isles. A large number of them settled in northern and eastern England in what would become known as the Danelaw, an area stretching from Yorkshire and Lancashire to East Anglia and the East Midlands. Since the Danelaw was essentially the land of the Angles, who were also from Denmark, it is not surprising that this region also has the highest incidence of blond hair in the British Isles today.
A lot has been written about the Anglo-Saxon conquest and historians have long debated the real genetic impact that the Anglo-Saxon had on the British population. Were the Romans and ancient Britons completely wiped out by the invaders or forced to retreat by to Wales, Cornwall and Scotland? Did the most of the native population intermingle with Anglo-Saxons creating a new hybrid ethnic group? What happened actually lies in between.
Germanic people brought a whole new set of paternal lineages with them, namely I1, I2a2a-Z161, R1a (L664 and Z284), R1b-U106, and to a lower extent Q1a. Those haplogroups now make up over half of all male lineages in England and Lowland Scotland.
One could expect to find an east-west gradient within England for Y-chromosomal haplogroups, since invaders came from the east and progressively advanced westward while intermingling with the indigenous population. Yet, overall England appears to be fairly homogeneous in terms of Germanic ancestry, except for Cornwall, which is slightly more Celtic. The only clear rift in Germanic ancestry is observed between England and Wales, and even actually within Wales, with western counties like Pembrokshire and Gwynedd having more Brythonic haplogroups (R1b-L21) and less Germanic ones. But it would be wrong to think of England as overwhelmingly more Germanic than Wales. Lineages seem to have evened out pretty well after 1,500 years. Nowadays, most English counties have between 55% and 65% of Germanic Y-DNA haplogroups, while Cornwall has 45% and Wales has much as 25%. The gap is not huge, but it clearly exists.
Nevertheless, an east-west gradient does exist between Gaulish Celtic or Roman lineages (both R1b-U152) in the east and Insular Celtic (R1b-L21) in the west. Cornwall and Wales have hardly any U152 at all, but boast respectively 40% and 65% of R1b-L21. If we exclude Germanic lineages, Ireland is almost purely Insular Celtic, L21 superseding U152 by a ratio to 50 to 1. The highest frequencies of Gaulish and Roman U152 are observed, unsurprisingly, closest to the continent, in southeast England and East Anglia.
While Danish Vikings settled mostly in eastern England, Norwegian Vikings colonised Orkney, the Shetland Islands, the northern and western coasts of Scotland, the Isle of Man, and in many parts of coastal Ireland, especially in Leinster and Munster. Norwegian Vikings carried a considerably higher percentage of haplogroup R1a-Z284, which is divided in two branches L448 and Z287. It is from this L448 branch that descended Somerled of Argyll (1100-1164), a military and political leader of the Scottish Isles who is the paternal ancestor of the members of Clan MacDougall and Clan Donald. The L176.1 subclade of L448 has to date been found almost exclusively among the descendants of Somerled. In contrast, the other Germanic branch of R1a, L664, is characteristic of West Germanic people like the Anglo-Saxons and is the most common variety in southern England.
Haplogroup Q represents a minor but distinctive Scandinavian lineage. It is found especially in the Danelaw, Orkney and Shetland, but also around Scotland, Lancashire, southern England, and, tellingly, Normandy. The Q1a2b1 (L527) subclade in particular is found almost exclusively in Scandinavia and in places settled by the Vikings. Q1a2a1a2 (L804) is also found in West Germanic countries and may have been brought by the Anglo-Saxons. Note that Q1b1a (L245) has also be found at trace frequencies in the British Isles, but may not be of Germanic origin. L245 is thought to have spread to Europe from the Middle East via the Phoenicians and the Jews, among others.
Normans
In the early 10th century, a Viking contingent made up of Danes and Norwegians, NorseGaels and Anglo-Danes sailed up the Seine River and laid siege on Paris. In exchange for not sacking the city, the King of France granted them land in northwest France if Rollo, the leader of the Norsemen accepted to become his vassal. This was born the Duchy of Normandy. The Norsemen adopted Christianity and the local Gallo-Romance language, from which evolved Anglo-Norman. They mixed extensively with the local French population and even accepted Frenchmen as part of their nobility.
In 1066, Duke William II of Normandy conquered England killing King Harold II at the Battle of Hastings. The invading Normans and their descendants replaced the Anglo-Saxons as the ruling class of England. The Anglo-Normans continued their expansion to Scotland (from 1072), Ireland (from 1169) and Wales (from the late 1200's). Thanks to their status and wealth, Norman noblemen left a greater number of offspring than the commoners, and as a result managed to leave a noticeable genetic impact.
For example, Norman surnames now make up 10% of all modern Irish surnames and even exceed 30% of all patronyms in Counties Wexford and Waterford, where the Normans first settled in the 12th century. Genetic genealogy tests have confirmed that Germanic Y-DNA haplogroups in the region are usually linked to Norman surnames (e.g. I1 for Powers, R1b-U106 for Roche and Sinclair). The reverse is not always true though, as some Irishmen took Norman names for social reasons. Additionally non-paternity events might have occured in some lineages over time, like everywhere else.
In Scotland, the Nomans founded several clans, known as the Scoto-Norman clans (Bruce, Comyn, Crawford, Fraser, Grant, Hall, Hay, Hunter, Innes, Jardine, Kerr, Menzies, Montgomery, Murray, Ramsay, Sinclair, Stewart, Sutherland), many of which would rise to the Scottish peerage, and some even provide future kings (Robert the Bruce, House of Stewart).
Distribution maps of Y-DNA haplogroups in Britain and Ireland

Y-DNA frequencies by region
| Region/Haplogroup |
I1
| I2a1
| I2a2
| R1a
| R1b
| G
| J2
| J1
| E1b
| T
| Q
| Sample size |
| England |
14 |
2.5 |
4.5 |
4.5 |
67 |
1.5 |
3.5 |
0 |
2 |
0.5 |
0.5 |
> 5000 |
| Cheshire & Lancashire |
15.5 |
0 |
7.5 |
2.5 |
69.5 |
0.5 |
1 |
0 |
2.5 |
0.5 |
0.5 |
301 |
| Cornwall & Devon |
8 |
1.5 |
3.5 |
4 |
75.5 |
2.5 |
2 |
0.5 |
1.5 |
0.5 |
0 |
375 |
| Cumbria |
12 |
1 |
2.5 |
6 |
70 |
0 |
1.5 |
0 |
6 |
0 |
0 |
117 |
| East Anglia |
19.5 |
2.5 |
8 |
5 |
56.5 |
4 |
1 |
0.5 |
3 |
0 |
0 |
466 |
| East Midlands |
16 |
1 |
5 |
4 |
63 |
1 |
4.5 |
1 |
4.5 |
0 |
0 |
200 |
| Home Counties |
10.5 |
1 |
4.5 |
3.5 |
68 |
1.5 |
2.5 |
0.5 |
3.5 |
0.5 |
0.5 |
420 |
| Northeast England |
14.5 |
0.5 |
4 |
5 |
65 |
3 |
2 |
1 |
1.5 |
0 |
1 |
286 |
| Southwest England |
15 |
1.5 |
4 |
5 |
72 |
2 |
3 |
0.5 |
3 |
0.5 |
0.5 |
357 |
| West Midlands |
17.5 |
0.5 |
4 |
3 |
66 |
1.5 |
2.5 |
0 |
3 |
0 |
0 |
190 |
| Ireland |
6 |
1 |
5 |
2.5 |
81 |
1 |
1 |
0 |
2 |
0 |
0 |
> 5000 |
| Ulster |
9 |
0.5 |
10 |
1.5 |
76.5 |
0 |
1.5 |
0 |
0 |
0 |
0.5 |
140 |
| Scotland |
9 |
1 |
4 |
8.5 |
72.5 |
0.5 |
2 |
0 |
1.5 |
0.5 |
0.5 |
> 5000 |
| Northeast Scotland |
13 |
4 |
6.5 |
4 |
69.5 |
0 |
2.5 |
0 |
0 |
0 |
0 |
76 |
| Southern Scotland |
19.5 |
1.5 |
11 |
3.5 |
57 |
1.5 |
1.5 |
1 |
3.5 |
0 |
0 |
118 |
| Western Scotland |
8.5 |
2.5 |
6.5 |
7.5 |
72 |
0 |
0.5 |
0 |
1.5 |
0 |
0 |
183 |
| Orkney |
10.5 |
1 |
2 |
22.5 |
62 |
0 |
0 |
0 |
0 |
1 |
0 |
159 |
| Shetland |
13.5 |
0 |
0.5 |
15.5 |
68.5 |
0 |
0 |
0 |
0 |
0.5 |
1 |
197 |
| Wales |
12 |
1 |
3 |
1 |
74 |
2.5 |
0.5 |
0 |
4 |
1 |
0 |
411 |
- East Anglia includes data for Cambridgeshire, Norfolk and Suffolk
- East Midlands includes data for Derbyshire, Leicestershire, Lincolnshire, Northamptonshire, Nottinghamshire and Rutland
- Home Counties includes data for Berkshire, Buckinghamshire, Essex, Kent, Middlesex, Surrey and Sussex
- Northeast England includes data for Durham, Northumberland and Yorkshire
- Southwest England includes data for Dorset, Gloucestershire, Hampshire, Oxfordshire, Somerset and Wiltshire
- West Midlands includes data for Shropshire, Staffordshire, Warwickshire and Worcestershire
Sources of Y-DNA frequencies